1. Introduction
Diatoms are an enormous group of microalgae containing more than 100,000 living species [1, 2]. A lot of work has been done to document diversity and relationships within the group. As a result, species-level taxonomy has become adequate to support the extensive and successful use of diatoms in biomonitoring based on their species-specific response to their environmental changes, especially organic pollution and eutrophication, with a broad spectrum of tolerance, from oligotrophic to eutrophic conditions [3, 4, 5].
Diatom taxonomy is based almost exclusively on the morphological features of the silica frustules [6] and, even after training and long experience, diatomists often find it difficult to agree about identifications. Moreover, morphological treatment is a time-consuming process and costly. In contrast, DNA-based identification has been proposed as an alternative method to complement or even replace traditional identification methods of species [7]. DNA barcoding is based on DNA sequences that are linked to morphologically identified specimens. It is considered a faster and universally applicable approach and also has the potential for refined analyses [8, 9]. Consequently, DNA barcoding deserves careful consideration as a means of improving the reliability of identifications and discovering species and also of increasing the quality and quantity of other taxonomic information. DNA-based methods are being used for assessing biodiversity in environmental samples, the so-called DNA metabarcoding [10], which allows one to determine a species’ presence and abundance from bulk samples of soil, water, or air [11]. Several studies have demonstrated that metabarcoding may be an applicable tool for ecological monitoring based on epilithic diatoms [12, 13, 14, 15, 16]. The preferred barcoding markers in these studies were 18S rRNA and rbcL (Ribulose-1,5-bisphosphate carboxylase/oxygenase).
The basis for a reliable identification with molecular methods is a comprehensive reference library where molecular and morphological data are tied together with a taxonomic name. However, the taxonomic consistency of the sequence information in these databases is very low for diatoms [17]. Rimet et al. [18] developed the R-syst::diatom barcoding library in open-access (http://www.rsyst.inra.fr/) that includes barcodes for microalgae strains of the culture collection maintained in INRA (Institut National de la Recherche Agronomique, France). Another limitation to DNA-based identification approaches is the natural intraspecific divergence of the barcoding marker in specimens from different geographic origins. To date, no information on barcoding sequences has been reported for diatom species from Ecuador. This is a constraint on the application of these methods in freshwater diversity studies and in biomonitoring.
In this context, this research aimed at obtaining the barcodes of Ecuadorian epilithic diatoms with a wide geographical distribution, a well-defined ecological range and characteristics that allow them to be reliable indicator species [19, 20, 21]. This study is a humble first step towards coupling the morphological and molecular data of benthic diatoms in Ecuador. We have therefore studied morphology as a diagnostic feature of each unialgal diatom strain via light and scanning electron microscopies and also determined the DNA barcode 18SV4 rRNA [9, 22] as well as rbcL [23, 24] in order to establish the foundation of a reference library. The original strains are kept alive in the Universidad de las Américas’ epilithic diatoms collection. By publishing these data online, we are contributing to a morphological and molecular taxonomic reference library for neotropical diatoms, making them available for comparative studies. Thus, we expect this to be a key step in developing a trophic water quality index based on a molecular biology analysis of epilithic diatoms in Ecuador.
2. Materials and methods
2.1. Sample collection
Samples were collected under the MAE-DNB-CM-2018-093 authorization conceded by the Environment Ministry of Ecuador. Epilithic diatom samples were collected from four freshwater streams within the Andean region of Ecuador (Figure 1; Table 1). Three of them characterized as páramo (highland wilderness) landscapes at the Iliniza Volcano (SP), Chimborazo volcano (CH) and Cajas national park (CA). The fourth one is located at the Jambelí stream, in the parish of Machachi, surrounded by agricultural livestock mosaics (Table 1). Location data were taken with a GPS navigator (WGS 84 UTM Zone 17 with 5 meters of precision). The sampling design aimed to capture indicator species from pristine streams (SP, CH and CA) and a eutrophic stream (JA) from the Ecuadorian Andean region (Figure 1).
Location of sampling sites along Ecuadorian Andes. JA: Jambeli River at Pichincha province. SP: San Pedro River at Illiniza Ecological Reserve. CH: stream at Chimborazo Ecological Reserve. CA: stream at Cajas National Park.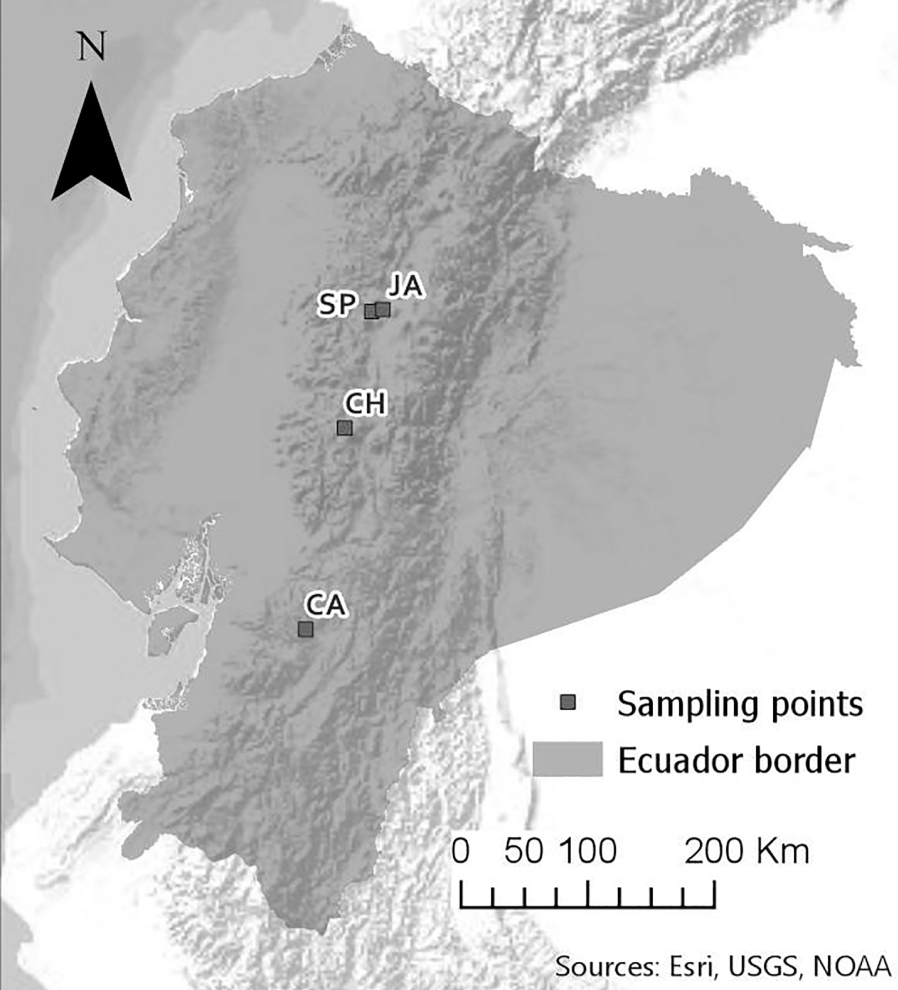
Location of the sampled streams in the central Ecuadorian Andean Region with geographical coordinates, altitude (masl) and the code assigned to samples collected in each stream
| Code | Province | Description | Latitude | Longitude | Altitude |
|---|---|---|---|---|---|
| SP | Pichincha | Illiniza Ecological Reserve | −0.58661 | −78.67343 | 3601 masl |
| JA | Pichincha | Quito | −0.57296 | −78.59418 | 3162 masl |
| CH | Chimborazo | Chimborazo Ecological Reserve | −1.41337 | −78.86458 | 3978 masl |
| CA | Azuay | Cajas National Park | −2.84271 | −79.14182 | 3165 masl |
The physical and chemical variables analyzed were water temperature, pH, turbidity, dissolved oxygen (DO) and conductivity, measured in the field with a multiparameter analyzer, and nitrate and phosphate, measured in the laboratory. Diatom samples were scrubbed off the upper surface of three to five submerged stones, 10 to 20 cm in diameter, using a toothbrush [25]. Diatom and water samples were kept cold until they were transferred to the laboratory.
2.2. Isolation and culturing
The environmental samples were cultured on BBM agar plates [26] supplemented with sodium silicate (10 mM) at 15–18 °C, light intensity of 50 μEm-2 s-1 and 12/12 hr light/dark regime. Individualized macroscopic colonies on agar plates were inoculated into liquid media and grown until they turned brown. This process was repeated until unialgal cultures were obtained. The achievement of unialgal cultures was checked by light microscopy using an Olympus BX51 microscope. Diatom strains were maintained on agar plates and liquid medium.
2.3. Morphology-based analysis
One fraction of each unialgal culture was used for morphological analysis. Living cells as well as cleaned frustules were examined and photographed by light microscopy. To remove organic material, the cells were oxidized with 50% nitric acid at 80 °C for an hour and rinsed several times with H2O [27]. Permanent slides were mounted with the high refraction index mounting medium Naphrax®. For scanning with the electron microscope, a few drops were dried on stubs and sputter-coated with gold using a Quorum Tech Q150RES. SEM images were taken using an ultra-high-resolution analytical field emission (FE) scanning electron microscope Tescan MIRA3, operated at 10 kV.
Species identification was performed using a microscope Olympus BX-40 according to the following taxonomic references [4, 28, 29, 30].
Sequence identity of the closest relative of each isolated strain found in GenBank using BLASTn
| Strain | 18SV4 | % identity | N hits | Closest match | rbcL | % identity | N hits | Closest match |
|---|---|---|---|---|---|---|---|---|
| CA01a | Nitzschia palea | 99–100% | 37 | KU341755.1 | Nitzschia palea | 98–100% | 104 | HF675122 |
| CH02a | Eolimna minima | 95–100% | 7 | KM084877.1 | Eolimna minima | 95–97% | 3 | KM084939.1 |
| JA01a | Achnanthidium minutissimum | 93–100% | 24 | KY863464 | Achnanthidium minutissimum | 97–100% | 15 | KY863482.1 |
| JA01b | Sellaphora seminulum | 93–100% | 6 | KR150677.1 | Sellaphora seminulum | 95–100% | 4 | KM084937.1 |
| JA01c | Sellaphora seminulum | 93–100% | 5 | KR150677.1 | Sellaphora seminulum | 95–100% | 4 | KM084937.1 |
| SP02a | Nitzschia fonticola | 90–98% | 3 | AJ867022.1 | Nitzschia fonticola | 99–100% | 4 | HF675068.1 |
2.4. Molecular-based analysis
A second fraction of each unialgal culture was used to extract total genomic DNA. Cells were pelleted by centrifugation of 1 mL culture medium at 8000 g for 2 min and disrupted using ceramic beads by vortexing. The extraction method was then performed according to Edwards et al. [31]. The rbcL and 18SV4 rRNA regions were submitted to PCR amplification using GoTaq® Green Master Mix by Promega with 0.2 μM of each primer. The V4 region of the 18S rRNA was amplified with the primer pair DIV4for/DIV4rev3 [13, 32]. Primer used for the rbcL fragment were Diat_rbcL_708F [33] and reverse primer R3 [34]. The selected primer pairs amplify a fragment around 300 pb and have been used previously for DNA metabarcoding analysis [12, 14, 24]. PCR regime included an initial denaturation at 95 °C for 2 min, then 35 cycles of denaturation at 95 °C for 45 s, annealing at 50 °C for 45 s, elongation at 72 °C for 30 s, and a final elongation at 72 °C for 2 min. Sequence reactions were performed with an Applied Biosystems® 3130 Genetic Analyzer. Sequences were edited with MEGA 7 software [35] and blasted against the GenBank database at NCBI (National Center for Biotechnology Information). Sequences of strains of the studied species from different geographical locations were downloaded from GenBank (Supplementary Tables S1 and S2) and aligned with barcoding sequences from Ecuadorian strains using the Muscle Algorithm [36]. Bolidomonas pacifica was used as an outgroup taxon, due to its genetic proximity to diatoms [37]. The ends of the alignments were trimmed to minimize missing characters. Tree topologies and branch lengths were computed separately for the two markers with the maximum-likelihood method (ML) using Tamura-Nei distance [38] with gamma distributed rates among sites followed by a statistical test of the tree topologies with 500 bootstrap replications.
3. Results
Five suitable bioindicator species were sampled from the Ecuadorian Andean Region (Table 1): Sellaphora seminulum (Grunow) D. G. Mann (strain JA01b, c), Nitzschia fonticola (Grunow) Grunow (strain SP02a) and N. palea (Kützing) W. Smith (strain CA01a) are tolerant to eutrophication; Eolimna minima (Grunow) Lange-Bertalot (strain CH02a) is a mesotrophic water bioindicator and Achnanthidium minutissimum (Kützing) Czarnecki (strain JA01a) is an oligotrophic water bioindicator. The comparison with the GenBank database of the barcoding regions obtained supported the morphological identification. Blasting results against NCBI are shown in Table 2. N. palea had the most hits (37 sequences for 18SV4; 104 for rbcL) followed by A. minutissimum (24 sequences for 18SV4; 15 for rbcL). GenBank accession numbers for 18SV4 rRNA and rbcL sequences from this study are MN589666-MN589670 and MN603956-MN603960 respectively.
JA01b strain presents linear valves with rounded apices. The axial area is narrow, linear. The central area is wide, rectangular. The raphe is filiform with the terminal fissures curved to the same side of the valve with distinct central pores. Striae are radiate, multiseriate, and more distant and curved near the central area, with 2 or 3 shorter striae. The areolas are rounded. Valve dimensions: Length: 9–12 μm; Width: 3–3.5 m, 20–22 striae in 10 μm (Figure 2i–l). This result is consistent with S. seminulum description [39]. Morphological description of JA01c strain also matches S. seminulum although its length ranges from 6 to 10 μm and width reaches up to 4.5 μm (Figure 2m–p). Thus, JA01c strain presents elliptical valve shape while JA01b has linear valves. Despite the morphological differences, both strains have 100% identity for 18SV4 and rbcL sequences. The number of 18S and rbcL sequences for Sellaphora seminulum in the GenBank database is scarce. There are strains from the Geneva basin [32] Seoul and Reunion Island [40] with a homology of 98% with 18SV4 and rbcL. Evans et al. [41] and Kermarrec et al. [24] described two strains of S. seminulum from United Kingdom and Mayotte Island at the Komora Archipelago, with a homology below 95%. There is a third case in the GenBank for a strain from Berlin that presents 100% of identity with rbcl, but less than 95% with 18SV4 [17].
Isolated diatoms under SEM: a–d: strain CA01a (N. palea (Kützing) W. Smith); e–h: strain SP02 (Nitzschia fonticola (Grunow) Grunow); a, i–l: strain JA01b (Sellaphora seminulum (Grunow) D. G. Mann); m–p: JA01c (Sellaphora seminulum (Grunow) D. G. Mann); q–t, u: JA01a (Sellaphora seminulum (Grunow) D. G. Mann); v–y: CH02a (Eolimna minima (Grunow) Lange-Bertalot). Scale bar = 1 μm.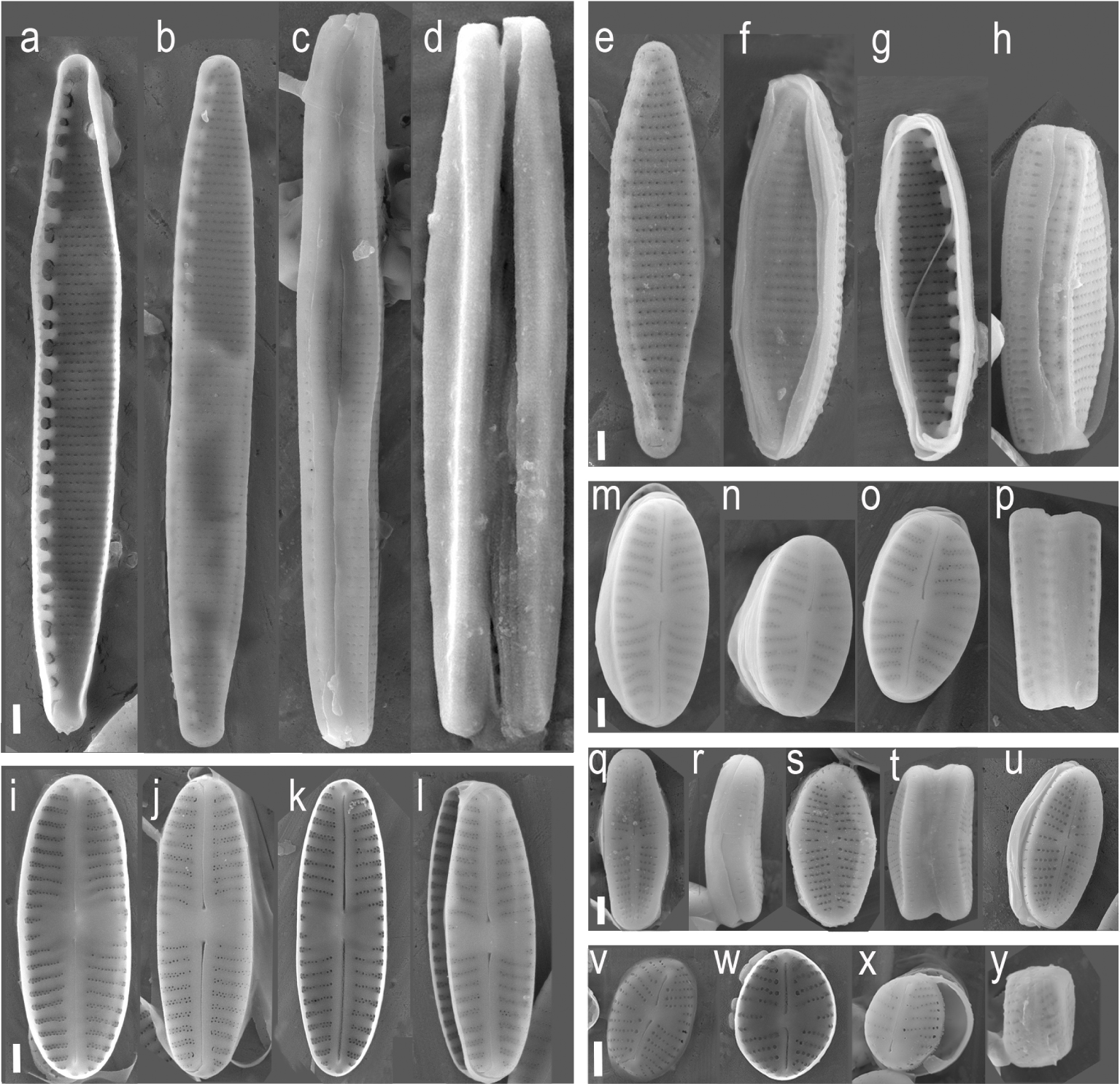
The morphological traits of strain SP02 match the species Nitzschia fonticola [42]: linear-lanceolate valves, acutely rounded apices, eccentric, marginal raphe, uniseriate striae on the valvar face and biseriate in the canal of the raphe, practically equidistant fibulae, except in the central area that presents two fibulae more distant from the others. Valve dimensions: Lenght: 15–18 μm; width: 2.5–3 μm; 10–14 fibulae in 10 μm (Figure 2e–h). The sequences obtained from this strain have higher identity with rbcL sequences (99–100%) than with 18SV4 sequences (90–98%) uploaded to the GenBank. There is no strain described as N. fonticola with both rbcL an 18SV4 barcodes reported. The sequences matching rbcL belong to strains collected from the United Kingdom, Spain, and Reunion Island [17, 40] while 18SV4 matches only one sequence (from France, data not published) with over 98% similarity. There are two other sequences for 18SV4 with less than 90% similarity from microorganisms isolated from karstic-stream biofilms in Germany (data unpublished).
CA01a strain, described as Nitzschia palea [43], presents linear-lanceolate valves, rostral apices, eccentric raphe, marginal, very thin uniseriate striae, equidistant fibulae including the central pair. Valve dimensions: Lenght: 25–27 μm; width: 2.8–3 μm; 16 fibulae in 10 μm (Figure 2a–d). There are plenty of sequences for N. palea [32, 37, 44] in the GenBank from all around the world, most of them with over 99% identity with CA01a strain for both rbcL and 18SV4 barcodes.
CH02a strain shows elliptical valves with rounded apices. The axial area is narrow, linear. The central area is little expanded, slightly rectangular, limited by the irregular size of the median striae. The raphe is filiform, straight, with proximal ends slightly flexed. The central pores are small. Striae are radiate, uniseriate, with irregular sizes near the central area, composed of rounded areolas (Figure 2v–y). Valve dimensions: Length: 6–10 μm; Width: 3–4 μm; 25–26 stretch marks in 10 μm. This is consistent with the morphological description of E. minima [45]. The closest match with both barcodings belongs to the Eolimna genera, but it is not identified at a species level [17]. There are other strains reported with a 99% identity match for 18SV4 described as Eolimna, Sellaphora and Navicula minima, which are homotypic synonyms. The rbcL sequence obtained showed 96% similarity with a strain identified as E. minima from Germany [17].
JA01a morphological traits are consistent with Achnantidium minutissimum [46]. It shows linear-lanceolate valves, rounded and rostrate ends, narrow axial area, linear, widening towards the center, straight raphe, filiform, curved terminal fissures, radiate striae, more widely spaced in the central area of the valve. Valve dimensions: Length: 6–7 m; Width: 2–3 m; 30–32 striae in 10 m. (Figure 2q–u). Blast results for both barcodes show from 98 to 100% identity with A. minutissimum from Europe, Asia and North America [9, 32, 37, 40]. There are a few strains from Japan and Île de Reunion that have less than 96% identity for 18SV4 (Data unpublished).
Phylogenetic trees were constructed based on 18SV4 and rbcL regions separately due to the lack of strains with both barcode sequences deposited in INSDC databases. The phylogenetic analyses include sequence representatives of each species from different geographical locations reported previously. In the phylogeny, both barcode sequences of the same species clustered into one clade (Figures 3 and 4). Species level clades are well supported statistically. Short branches in species clades mean low divergence between the Ecuadorian strains and their homologous strains. In the phylogenetic tree based on the 18SV4 region (Figure 3), N. fonticola sequences are in the same clade as E. minima and S. seminulum but separated from N. palea. However bifurcations representing the relationship between the species are not well supported by bootstrap values. Nevertheless, in the phylogeny of rbcL (Figure 4), sequences belonging to Nitzschia genera clustered together. E. minima and S. seminulum are grouped in the same clade in both phylogenetic analyses.
Phylogram constructed using 18SV4 rRNA sequences. The phylogram was constructed using the maximum-likelihood method with Tamura-Nei protocol. Numerical values at the nodes of the branches indicate bootstrap values above 50%. Accession number and geographical origin is indicated for each sequence (Supplementary Table S1).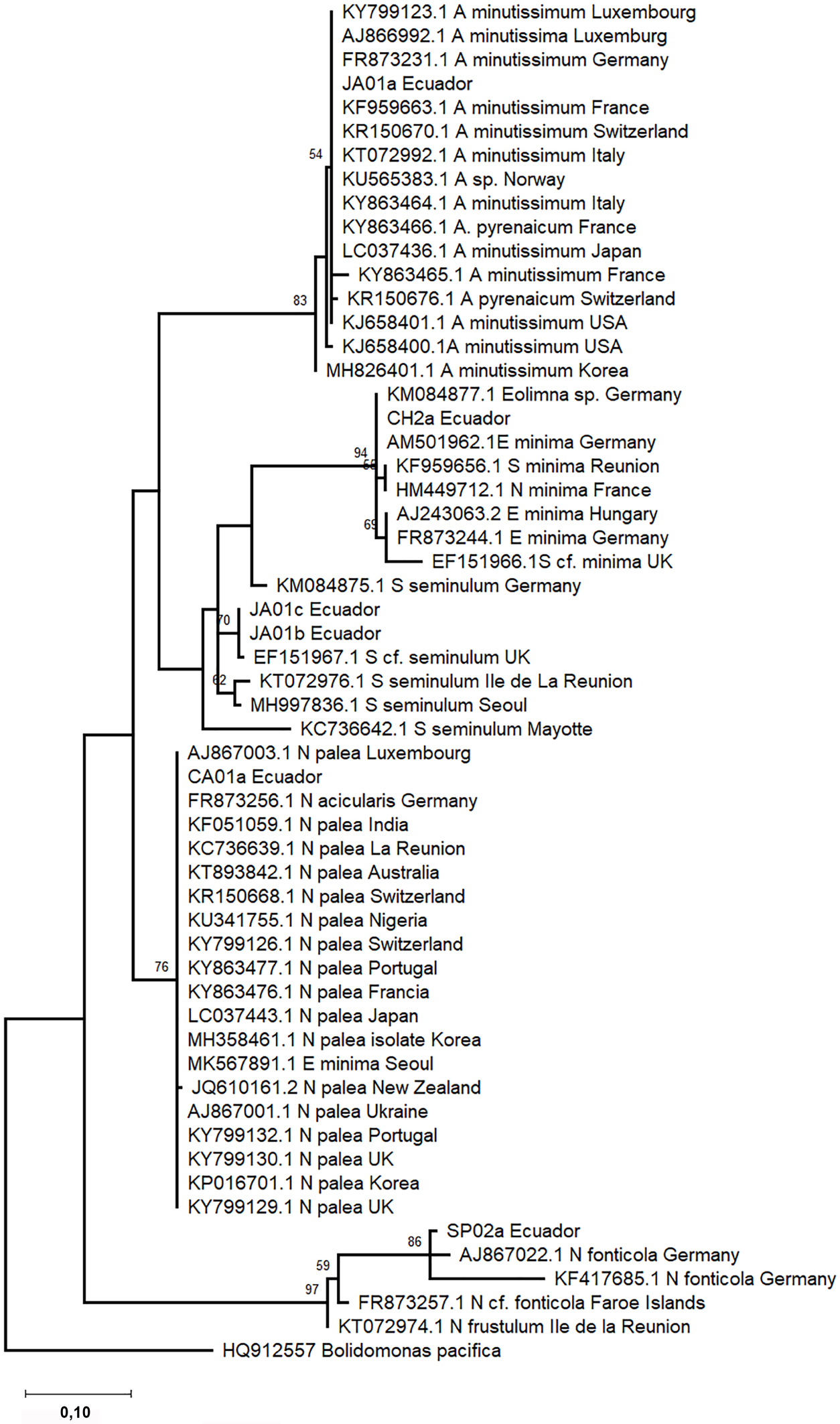
Phylogram constructed using rbcL sequences. The phylogram was constructed using the maximum-likelihood method with Tamura-Nei protocol. Numerical values at the nodes of the branches indicate bootstrap values above 50%. Accession number and geographical origin is indicated for each sequence (Supplementary Table S2).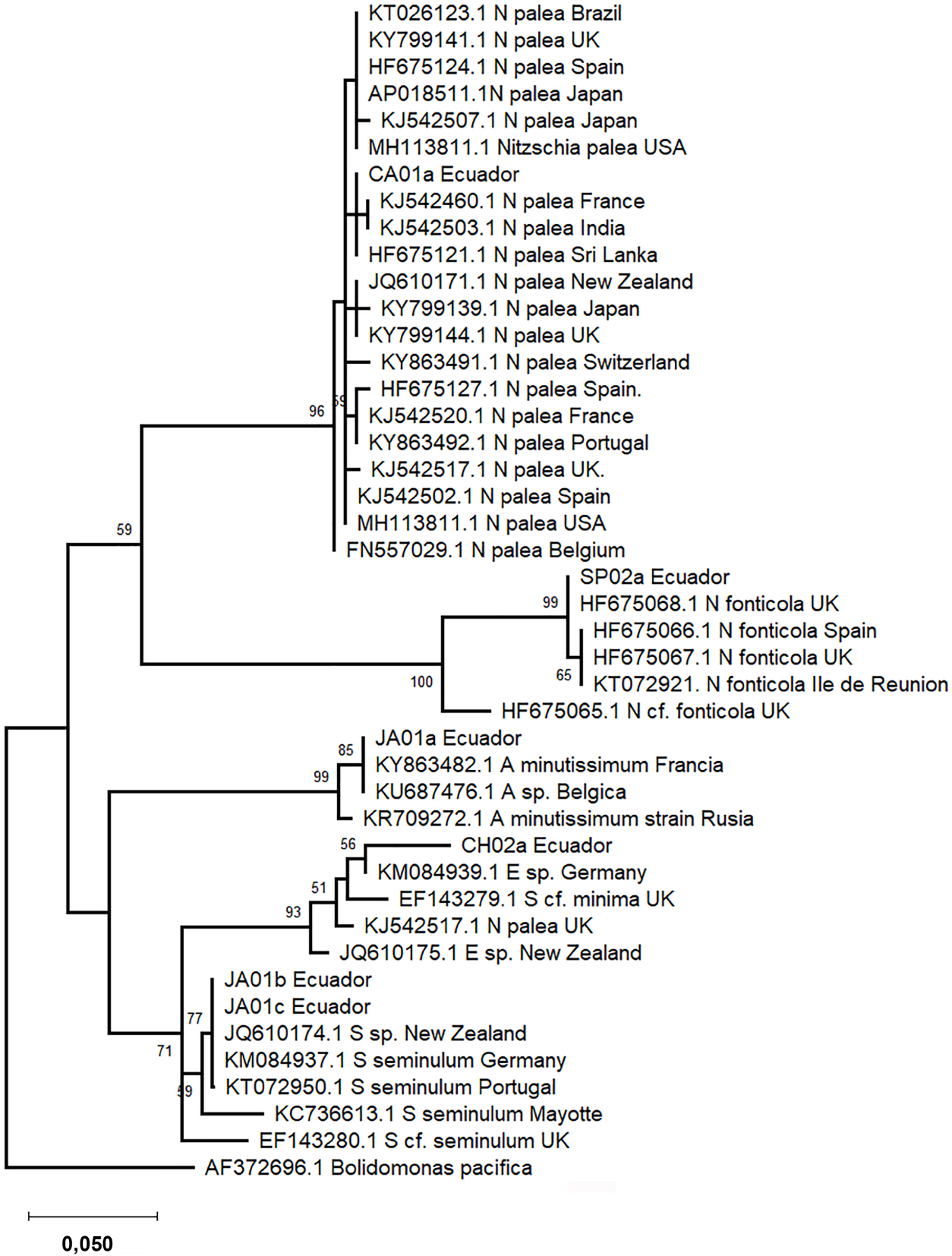
4. Discussion
To the best of our knowledge, this study is the first attempt to establish a barcode reference library for epilithic diatoms from Andean streams of Ecuador. Five strains considered as water quality bioindicators have been cultured and sequenced for rbcL and 18SV4 barcodes. N. palea, N. fonticola and S. seminulum are tolerant to organic pollution and eutrophication. N. palea regularly occurs downstream of sewage treatment plants [47]. E. minima is abundant in mesotrophic water. A. seminulum is a small monoraphid diatoms of oligo/mesosaprobic rivers [47]. These species are very important from an ecological point of view, since they are often among the most abundant benthic species in freshwater systems. Furthermore, N. palea, A. minutissimum and E. minima have been used to assess sensitivity to herbicides [37, 48] and metals [49].
The occurrence and/or abundance of epilithic diatoms to define ecological status through morphological characteristics has some limitations [7]. The use of DNA sequences could potentially alleviate these limitations [50]. The barcoding sequences of the strains presented here have a high percentage of identity with the sequences reported in INSDC databases for the same species. Hence, DNA barcoding, based on 18SV4 and rbcL, emerges as a useful tool for diatom species identification in Andean streams. Nonetheless, we found that there are reported sequences for the same species with a divergence of at least 5%. This could be due to an inaccurate identification, since there are poor circumscriptions and a lack of reliable information about the epithets ‘minima’, and ‘seminulum’ even though they are apparently well–established and often referred to in ecological and taxonomic literature [51, 52, 53]. For instance, E. minima can barely be identified using light microscopy (LM) and the lack of illustrations from a scanning electron microscope (SEM) is a major impediment [7]. Moreover, there are identification problems derived from isolation and culturing, which are needed to obtain barcodes, given that some species experience cell size reduction in culture [54]. That could be the case for S. seminulum strains, JA01b and c, which are different in size but identical for rbcL and 18SV4 sequences. This supports the necessity of barcode reference libraries that couple morphological and molecular data for comparative analyses [12, 17].
The usefulness of 18SV4 and rbcL barcodes to assess phylogenetic relationships in diatoms has been proven extensively [6, 17, 18, 55, 56]. The topology of the phylogenetic trees demonstrates that epilithic diatoms from Ecuador show close relatedness to those of same species isolated from other geographical regions. This could mean low phylogeographic variability in diatoms. However, the E. minima clade presents the highest divergences, but this could be due to identification problems related with this species. The genera clades are congruent with previous analysis where Sellaphora falls into one group with Eolimna [17] and these are separate from Nitzschia and Achnantidium [18].
This research highlights the complementary aspects of classical taxonomy and DNA barcoding. All reference sequences presented here are linked to morphological detailed images in order to initiate a complete barcode reference library for diatoms in the neotropical region. In addition, this study demonstrates that it would be feasible to use the existing barcoding data for diatoms to develop molecular tools for bioassessment of aquatic ecosystems in the Ecuadorian Andean region.
Acknowledgements
This study was funded by the Universidad de Las Américas [grant number: BIO.IBR.19.05], Quito, Ecuador. Access to collection resources was granted by means of the Framework agreement for Access to genetic resources: MAE-DNB-CM-2018-0093 celebrated between the Environment Ministry, Ecuador and the Universidad de Las Américas, Ecuador. Thanks to Miguel Martínez-Fresneda Mestre for helping with the maps.
Supplementary data
Supporting information for this article is available on the journal’s website under https://doi.org/10.5802/crbiol.2 or from the author.





 CC-BY 4.0
CC-BY 4.0