1 Introduction
Recent advances in the molecular phylogenetics of stingrays (Myliobatoidei: [1]) have challenged the previously accepted taxonomy of species in a number of genera [2–6]. Both a cytochrome-oxidase I gene (COI)-based barcoding survey of Australasian chondrichthyans [7], a recent COI-barcoding survey of northwestern Australian myliobatoids [8] and a recent systematic survey of elasmobranchs based on the nicotinamide-adenine dinucleotide dehydrogenase subunit two gene marker [3] have pointed to problematic cases relative to the definition of species boundaries, when divergent haplotypes are observed within a given nominal species. One of these cases is the blue-spotted maskray, Neotrygon kuhlii (Müller and Henle, 1841) [9].
The blue-spotted maskray under its current taxonomic definition has a wide Indo-Pacific distribution, from the Red Sea to southern Africa and from the western Indian Ocean to the western Pacific, reaching Japan in the North, Tonga in the South-East and New Caledonia in the South [10]. It is currently the most frequently landed stingray species in the Coral Triangle [11], hence it is a species of commercial interest but also one of conservation concern given the general vulnerability of elasmobranchs to fishing pressure [12,13]. N. kuhlii shows unusually high within-species diversity, with average nucleotide distances among haplotypes reaching 2.8%–3.0% at the COI locus [7,8]. An intron-marker based diversity study has confirmed the remarkably high degree of genetic differentiation among populations, but has failed to detect multiple cryptic species within Coral Triangle N. kuhlii [14]. One study [8] reported 3.5% nucleotide divergence at the COI locus between a sample from the Great Barrier Reef and samples from northwestern Australia and the Coral Triangle. A recent phylogeographic survey of Neotrygon spp. [15] has confirmed this preliminary result. Blue-spotted maskray from the Coral-Sea deserve particular scrutiny as argued in the following.
The New Caledonian maskray, N. trigonoides Castelnau, 1873 [16] is currently considered as a junior synonym of N. kuhlii [15,17,18]. In his description, F. de Castelnau [16] noted a large number of small black spots on the dorsal surface of the stingray whereas in their description of N. kuhlii, J. Müller and J. Henle [9] only mentioned “einzene, kleine, runde blaue Flecken (3-6) auf jeder Brustflosse” and provided a figure of N. kuhlii that illustrates this description (Fig. 1A). The two species, N. kuhlii and N. trigonoides, were synonymized by P.R. Last and W.T. White [17] who compared the holotype of Castelnau's N. trigonoides to what they identified as N. kuhlii from eastern Australia: “A close examination of the holotype of Raya trigonoides Castelnau, 1873 from New Caledonia… confirmed that it is conspecific with eastern Australian forms of N. kuhlii”.
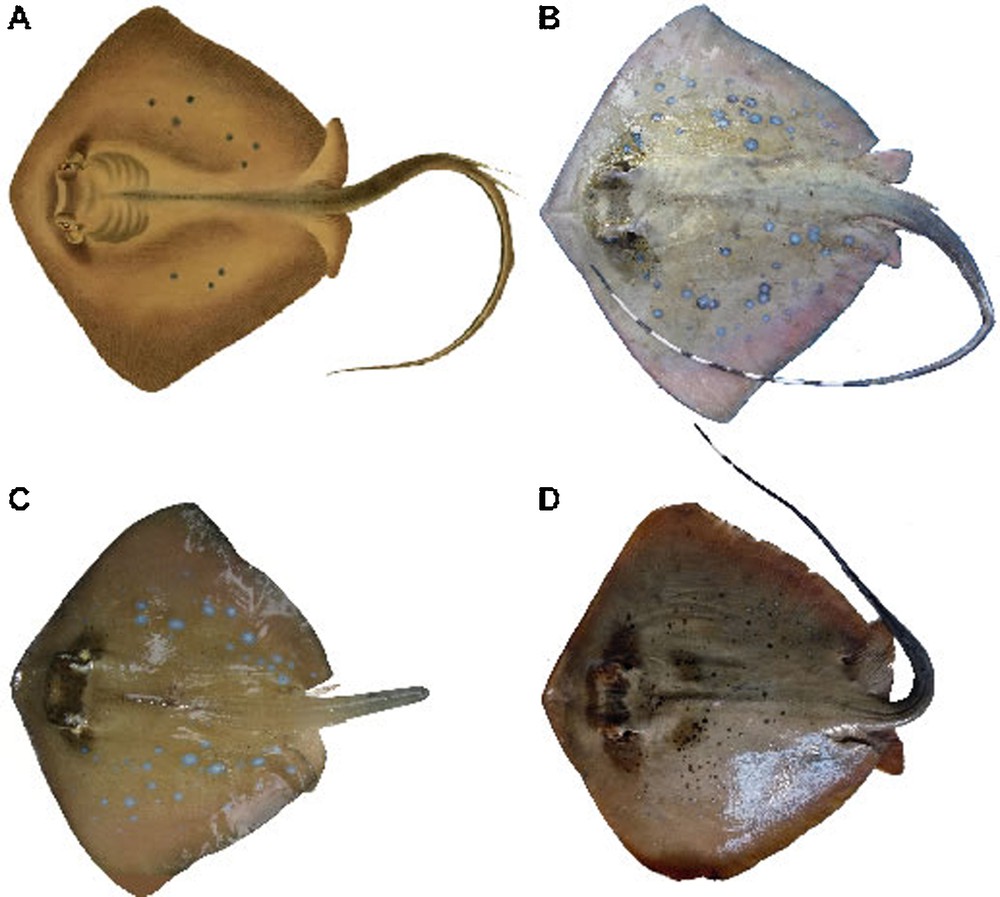

Specimens of blue-spotted maskrays from various locations showing variation in pigmentation patterns. A. Original drawing of a specimen (presumably type material) of Neotrygon kuhlii (pl. 51 in [9]). B. Specimen MZB 20843 (male N. kuhlii; no. 2 in Table 1) from Meulaboh, Aceh (04°07′N 96°08′E; 28 April 2009; I.S.A.). C. Specimen MZB 20851 (male N. kuhlii, 260 mm DW; no. 8 in Table 1) from Pulau Pari, Java Sea (05°51′S 106°37′E; 14 December 2008). D. Specimen MNHN 2009-0823 (female N. trigonoides, 350 mm DW; no. 38 in Table 1) from Saint-Vincent Bay, New Caledonia (21°57′S 166°02′E; 07 March 2009; P.B.).
In the present work, we re-examine spotting patterns in Castelnau (1873)’s N. trigonoides [16] and compare them to Müller and Henle (1841)’s N. kuhlii [9] and to blue-spotted maskray specimens from a range of locations in the Indo-West Pacific. We examine COI gene sequences for samples of blue-spotted maskray from two distant sites in the Coral-Sea, and from a number of sites in the Indian Ocean and in the Coral Triangle. Finally, we provide a redescription of N. trigonoides based on the COI gene sequence.
2 Materials and methods
2.1 Samples
Table 1 lists the blue-spotted maskray individuals sampled for DNA and barcoded for the present survey. This list includes 13 newly sampled specimens from the Indian Ocean (nos. 1–4 in Table 1), the Coral Triangle (nos. 7–9; 28; 30–32) and New Caledonia (nos. 37–39). All newly sampled specimens were deposited in zoological collections (Table 1) except no. 1 (from Pemba Island, Tanzania), which was not retained. Table 1 also includes references to sequence data retrieved from GenBank (http://www.ncbi.nlm.nih.gov/; accessed 25 November 2012) to which the new sequences were compared.
Specimens of blue-spotted maskray (Neotrygon kuhlii) and New Caledonian maskray (N. trigonoides) analysed for nucleotide sequence variation at the COI gene locus.
| Region (species), sampling location | Coordinates | Sampling date | N | Collector | Fig. 2 no. | Collection no. | Tissue no. | Photograph | GenBank accession no. |
| Indian Ocean (N. kuhlii) | |||||||||
| Pemba Island, Tanzania | 05°21′S 39°38′E | May 2010 | 1 | JDD | 1 | – | IRD zanz1 | b | KC295416 |
| Meulaboh, Aceh | 04°07′N 96°08′E | April 2009 | 1 | ISA | 2 | MZB 20843 (LIPI 4406) | ME3 | Fig. 1B | JX304805 |
| Perbaungan, Malacca Strait | 03°39′N 98°59′E | December 2009 | 1 | ISA | 3 | MZB 20847 (LIPI 4401) | MS_KL3 | b | JX304818 |
| Padang, Sumatera | 00°56′S 100°21′E | August 2009 | 1 | ISA | 4 | MZB 20845 (LIPI 4411) | PD1 | - | JX304828 |
| Ningaloo Reef | 21°55′-22°35′S | ||||||||
| 113°39′-113°53′E | August–September 2010 | 2 | O. O'Shea | 5, 6 | – | NKNR42, 43 | - | JQ765536–JQ765537 | |
| Coral Triangle (N. kuhlii) | |||||||||
| Pulau Pabelokan, Java Sea | 05°27′S 106°29′E | Aug. 2009 | 1 | ISA | 7 | MZB 20852 (LIPI 4410) | PB2 | b | JX304829 |
| Pulau Pari, Java Sea | 05°51′S 106°37′E | December 2008 | 1 | Mumu | 8 | MZB 20851 (LIPI 4402) | PR | Fig. 1C | JX304836 |
| Pulau Peniki, Java Sea | 05°46′S 106°38′E | March 2009 | 1 | ISA | 9 | MZB 20850 (LIPI 4399) | PN5 | b | JX304840 |
| Haiphong, Viet Nam | 20°46′N 106°52′E | September 2010 | 2 | - | 10, 11 | – | NKVN74, 75 | - | JQ765561, JQ765562 |
| Java Sea | - | April 2004 | 5 | W.T. White | 12–16 | – | BW A2575–BW A2579 | - | EU398737–EU398741 |
| Tanjung Manis, South China Sea | 02°07′N 111°19′E | April 2004 | 1 | J. Caira, K. Jensen, C. Healy | 17 | – | GN3636 = BO424 | b | JN184065 |
| South China Sea | 05°20′N 110°26′E | 2011 | 1 | - | 18 | – | FBBGC040-11 | - | JQ681494 |
| Bali | 08°45′S 115°10′E | August 2002 | 1 | W.T. White | 19 | CSIRO H 6124–01 | BW A2580 | - | EF609342 |
| Bali | 08°45′S 115°10′E | April 2004–March 2005 | 5 | W.T. White | 20–24 | – | BW A2571–BW A2574, BWA2583 | - | EU398736, EU398742–EU398745 |
| Bali | 08°44′S 115°11′E | January 2008 | 1 | PB | 25 | – | NK_BL | b | JX304860 |
| Penghu, Taiwan | ∼23°37′N ∼119°36′E | May 2005 | 3 | - | 26–28 | – | BW A2584–BW A2585 | - | EU398733–EU398735 |
| West coast, Taiwan | - | Oct. 2010 | 1 | H.C. Ho | 29 | – | wjc627 = 20101017HBH | - | JX304868 |
| Ishigaki-shima, Ryukyu Islands | ∼24°18′N ∼124°10′E | November 2004 | 1 | - | 30 | NSMT P–91858 | - | - | AB485685 |
| Ambon, Molucca Islands | 03°40′S 128°11′E | October 2008 | 1 | I.S. Arlyza, La Pay | 31 | MZB 20864 (LIPI 4400) | AM1 | b | JX304892 |
| Kei Island, Molucca Islands | ∼07°37′S ∼135°20′E | April 2009 | 1 | A. Kusnadi | 32 | MZB 20866 (LIPI 4405) | ARA1 | b | JX304898 |
| Biak, West Papua | 00°58′S 136°16′E | May 2009 | 1 | Alvi | 33 | MZB 20867 (LIPI 4408) | BK5 | b | JX304909 |
| Gulf of Carpentaria | 12°28′S 141°29′E | March 1995 | 1 | - | 34 | – | BW A208 | - | DQ108184 |
| Coral-Sea (N. trigonoides) | |||||||||
| Lizard I., Great Barrier Reef | 14°41′S 145°27′E | December 2008 | 3 | MGM | 35–37 | – | NKGBR39, 40; NKNR41 | - | JQ765534, JQ765535 |
| St Vincent Bay, New Caledonia | 21°56′S 165°55′E | August 2008 | 1 | P. Morlet | 38 | CSIRO uncat. a (NC 20080816) | Dkuh 20080816 (NC1) | b | JX304916 |
| St Vincent Bay, New Caledonia | 21°57′S 166°02′E | March 2009 | 1 | P. Morlet | 39 | MNHN 2009–0823 | Dkuh 20090307 (NC2) | Fig. 1D | JX263420 |
| St Vincent Bay, New Caledonia | 21°57′S 166°02′E | August 2008 | 1 | P. Morlet | 40 | IRDN 20090816 | Nkuh 20090816 (NC3) | b | JX304917 |
a Formalin/alcohol-preserved specimen (female, 22.5 cm disc width) sent by PB to Alistair Graham, CSIRO, Hobart, on 14 August 2009.
b Photograph posted in Supplementary Material, Fig. S1.
2.2 Nucleotide sequences
A tissue fragment ∼0.05 cm3 to ∼1 cm3 was removed, using surgical scissors, from the pelvic or pectoral fins, or the tail and was preserved in 95% ethanol at ambient temperature. DNA extraction was done using either the Viogene (Taiwan) tissue genomic DNA extraction protocol, or the DNEasy DNA extraction kit of Qiagen GmbH (Hilden, Germany). DNA was stored in 1X, pH 8.0 TE buffer (AppliChem, Darmstadt, Germany). Polymerase chain reaction (PCR) amplification of a fragment of the COI gene was done in a T-Gradient thermal cycler (Biometra, Göttingen, Germany) using 20 μL reaction mixture with the following concentrations: 0.05 units/μL Taq DNA polymerase in 2× DFS-Taq Mastermix (Bioron GmbH, Ludwigshafen, Germany), 16 mM (NH4)2SO4, 65 mM TrisHCl, pH 8.8 at 25 °C, 0.01% Tween-20, 2.75 mM MgCl2, 0.8 mM dNTP mix, 0.4 μM of each primer (AITBiotech, Singapore) and 2 μL DNA template. The primers were FishF1 (5′- T C A A C C A A C C A C A A A G A C A T T G G C A C -3′) and FishR1 (5′- T A G A C T T C T G G G T G G C C A A A G A A T C A -3′) [19]. PCR parameters were an initial denaturation at 94 °C for 2 min, followed by 35 cycles of heating (94 °C for 1 min), annealing (47 °C for 1 min) and extension (72 °C for 1 min) with a final extension step at 72 °C for 10 min.
Individual MNHN 2009-0823 was also PCR-amplified for the complete cytochrome b gene, using primers L14735 (5′- A A A A A C C A C C G T T G T T A T T C A A C T A -3′) and CB7 (5′- C T C C A G T C T T C G G C T T A C A A G -3′, slightly modified from CB6ThrH-15930) [20,21]. The reaction volume was 15 μL and the reaction mixture contained 0.2 mM dNTPs, 1.5 μL 10 × PCR buffer (Bioman, Taipei), 0.5 μM each of forward and reverse primer, 0.2 U Taq DNA polymerase (Bioman), and 1.0 μL template DNA. The PCR parameters were initial denaturation at 94 °C for 4 min followed by 35 cycles of denaturation (94 °C for 45 s), annealing (48 °C for 1 min), and extension (72 °C for 1 min), and a final extension step at 72 °C for 10 min.
PCR products were visualized on 1% agarose gels and their size was estimated as approximately 670 bp. After purification (by isopropanol precipitation), 1 μL 1/8-diluted PCR product was subjected to sequencing reaction, in both forward and reverse directions, using the BigDye Terminator v3.1 cycle-sequencing kit (Applied Biosystems, Foster City, CA, USA). Cycling conditions were according to the manufacturer's protocol. Sequencing reaction products were cleaned by removing dye-terminator (CleanSEQ kit, Beckman Coulter, Beverly, MA, USA) and loaded onto an ABI Prism 3100 DNA sequencer (Beckman Coulter).
2.3 Phylogenetic analysis
The phylogenetic analysis package Mega5 [22] was used to estimate nucleotide distance between major clades and nucleotide diversity within. Among the nucleotide substitution models proposed to this effect by Mega5, the most likely according to the Bayesian information criterion was the Tamura 3-parameter model [23] (T92) with non-uniform evolutionary rates among sites modelled by discrete gamma distribution (+G). Therefore, nucleotide distance and diversity were estimated according to the T92+G model.
Phylogenetic analyses were done using the Neighbour-joining (NJ) algorithm on T92+G-modelled genetic distances (MEGA5) and partitioned Maximum-likelihood (ML) as implemented in the RAxML-HPC [24] with its graphical interface raxmlGUI 0.93 [25]. For the partitioned ML search with the mixed model of nucleotide substitution, a GTR+G model (with four discrete rate categories) was used for each partition (respective to codon position) as RAxML only provides GTR [26] -related models of rate heterogeneity for nucleotide data [24]. ML tree search was done with 100 separate runs using the default algorithm of the program from a random starting tree for each run. The final tree was selected among suboptimal trees in each run by comparing likelihood scores under the GTR+G model. Nodal support was assessed by bootstrapping the original matrix of sequences [27] with the NJ algorithm and ML criterion, based on 1000 pseudo-replicates.
2.4 Spotting patterns
The sample of individuals chosen for studying spotting patterns included the holotype of N. trigonoides (NMMV 51684), the specimen represented Plate 51 of [9] (presumably one of the three syntypes of N. kuhlii), and those specimens analysed at the COI locus of which we possessed a photograph of quality suitable to counting and sizing all spots present on the dorsal surface of the disk (Fig. 1 and Supplementary material, Figs. S1 and S2).
The number of ocellated spots on the dorsal side of an individual was counted and their diameter was measured relative to disk width (DW). The maximal diameter of an ocellated spot was then assigned to one of three classes chosen arbitrarily: small, when diameter ≤ 2% DW; medium, when diameter was comprised between 2% and 4% DW; and large, when diameter > 4% DW. Dark-brown or black spots (> 1% DW) and speckles (≤ 1% DW) on the dorsal side of disk were also counted. Dark spots and speckles located in the mask area were not counted, because the quality of some of the pictures was not good enough to distinguish them from the brown background in this part of the head (Supplementary material, Figs. S1 and S2). The symmetrical pair of brown blotches in the scapular region was encoded 0 (when absent), 1 (when visible) or 2 (when conspicuous). Spotting patterns were compared among individuals through correspondence analysis (CA) [28]. CA was run using the FactoMineR package [29] under R [30]. Only dark spots and speckles are clearly visible on specimen NMV 51684. The ocellated white spots mentioned by Castelnau [16] have apparently faded in preservative, as noted previously [17]. Because its spotting patterns were characterized by three descriptor variables only (number of black speckles, number of black dots and presence/absence of brown scapular blotch), specimen NMV 51684 was included as active element in a second CA run based on these three descriptor variables.
3 Results
The NJ tree of COI haplotypes (Fig. 3) showed a main dichotomy separating maskrays from the Coral-Sea (northern Great Barrier Reef, New Caledonia) from all the other blue-spotted maskray sampled in the Indo-West Pacific. The net nucleotide divergence between the two clades was 0.026 whereas the nucleotide diversity within clades was 0.003 and 0.027, respectively. By comparison, the net nucleotide distance between N. leylandi and N. picta was 0.031. The ML tree constructed on the basis of the GTR+G model yielded a slightly different topology (not shown), where the Coral-Sea maskray haplotypes formed a haplogroup external to N. kuhlii, but whose monophyly was not supported enough statistically. In all other aspects, the ML tree was similar to the NJ tree. Bootstrap scores for the ML tree were added on the NJ tree (Fig. 3). Thus, both the NJ tree and ML tree of COI haplotypes recovered monophyly of the blue-spotted maskray, but yielded different results on whether the Coral-Sea maskrays formed a strongly supported monophyletic subgroup. In spite of the slightly incongruent results based on two tree reconstruction methods, both clearly separated maskrays from the Coral-Sea from all the other blue-spotted maskrays sampled in the Indo-West Pacific.
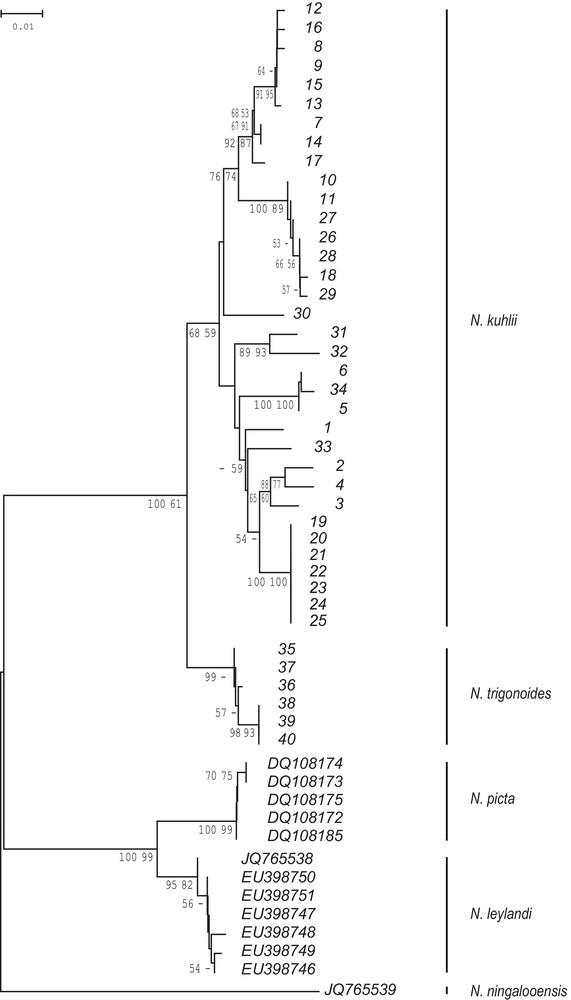
Neighbour-joining (NJ) tree of partial COI gene nucleotide sequences (T92+G-modelled nucleotide distances; Mega5 [22]) from 34 blue-spotted maskray, Neotrygon kuhlii, specimens collected distribution-wide and 6 New Caledonian maskrays, N. trigonoides, from the Coral-Sea. Tree rooted by homologous sequences from N. leylandi, N. ningalooensis, and N. picta. Distance scale bar, percent bootstrap values (> 50%; 1000 bootstrap resamplings; Mega5) are given. The first percentage indicated at a node is the bootstrap score relative to the NJ tree; the second percentage concerns the maximum-likelihood phylogeny constructed under RAxML (GTR+G model; [26]). Specimens numbered as in Table 1 and Fig. 2.
Spotting patterns of New Caledonian maskray specimens were markedly different from all other N. kuhlii specimens examined (Table 2). Diagnostic differences included the absence of medium or large ocellated spots in New Caledonian maskray, whereas all other N. kuhlii specimens possessed a substantial proportion of these; the presence of a few dark spots in all specimens from New Caledonia, vs. absence in all other N. kuhlii specimens except one; and the presence of a pair of brown scapular blotches, exclusively in specimens from New Caledonia. Moreover, ocellated spots were rather pale grey in New Caledonian maskray specimens vs. pale-blue to blue in all other N. kuhlii specimens (Supplementary material, Figs. S1 and S2). Pictures of blue-spotted maskray taken underwater at Lizard Island (Supplementary material, Fig. S2), which could not be utilized for the present morphological analysis, revealed features similar to those characterizing New Caledonian specimens, i.e. numerous dark speckles, a few dark spots, and the scapular brown blotch.
Neotrygon kuhlii and N. trigonoides. Matrix of individuals characterized by the numbers of ocellated spots [sorted into three size-classes: small: ≤ 2% disk width (DW); medium: > 2% DW and ≤ 4% DW; large: > 4% DW], the number of dark speckles (≤ 1% DW), the number of dark spots (> 1% DW), and the absence or presence of a scapular brown blotch (0: absent; 1: visible; 2: conspicuous) on the dorsal surface of left or right half-disk.
| Species, individual no. | Sampling region | Side of disk | N ocellated spots | N dark speckles | N dark spots | Scapular blotch | |||
| Fig. 2 no. | Specimen no. | Small | Medium | Large | |||||
| Neotrygon kuhlii | |||||||||
| - | Pl. 51 of [9] | Unknown | Left Right |
2 1 |
1 5 |
0 0 |
2 1 |
0 0 |
0 0 |
| 1 | zanz 1 | Tanzania | Left Right |
44 51 |
29 21 |
2 2 |
10 15 |
0 0 |
0 0 |
| 2 | MZB 20843 | Aceh | Left Right |
28 29 |
13 11 |
0 0 |
16 16 |
0 0 |
0 0 |
| 3 | MZB 20847 | Malacca Strait | Left Right |
2 4 |
14 10 |
0 1 |
5 3 |
0 0 |
0 0 |
| 7 | MZB 20852 | Java Sea | Left Right |
5 9 |
8 3 |
1 0 |
21 17 |
0 0 |
0 0 |
| 8 | MZB 20851 | Java Sea | Left Right |
22 18 |
9 6 |
2 2 |
12 8 |
0 0 |
0 0 |
| 9 | MZB 20850 | Java Sea | Left Right |
12 22 |
12 11 |
2 0 |
2 1 |
0 0 |
0 0 |
| 17 | BO424 | S. China Sea | Left Right |
35 41 |
22 6 |
0 0 |
16 13 |
0 0 |
0 0 |
| 25 | NK BL | Bali | Left Right |
21 24 |
6 5 |
0 0 |
43 54 |
0 0 |
0 0 |
| 31 | MZB 20864 | Ambon | Left Right |
9 2 |
7 12 |
0 0 |
5 2 |
0 0 |
0 0 |
| 32 | MZB 20866 | Kei Islands | Left Right |
36 30 |
9 9 |
0 0 |
53 53 |
0 1 |
0 0 |
| 33 | MZB 20867 | Biak | Left Right |
41 37 |
25 13 |
0 2 |
10 7 |
0 0 |
0 0 |
| Neotrygon trigonoides | |||||||||
| - | NMV 51684 | New Caledonia | Left Right |
ND ND |
ND ND |
ND ND |
24 26 |
4 2 |
1 1 |
| 38 | CSIRO uncat. | New Caledonia | Left Right |
54 38 |
0 0 |
0 0 |
36 25 |
1 1 |
2 2 |
| 39 | MNHN 2009-0823 | New Caledonia | Left Right |
33 28 |
0 0 |
0 0 |
85 91 |
10 8 |
2 2 |
| 40 | IRDN 20090816 | New Caledonia | Left Right |
15 16 |
0 0 |
0 0 |
55 57 |
13 13 |
2 2 |
Correspondence analysis of the matrix presented in Table 2 determined two main clusters (Fig. 4A, B). One cluster exclusively comprised the specimens from New Caledonia; the analysis placed the holotype of N. trigonoides within this cluster (Fig. 4B). The other cluster comprised all N. kuhlii specimens from the Indian Ocean and the Coral Triangle and also comprised the N. kuhlii specimen referred to by Müller and Henle [9] along their description of the species (Fig. 4A, B).
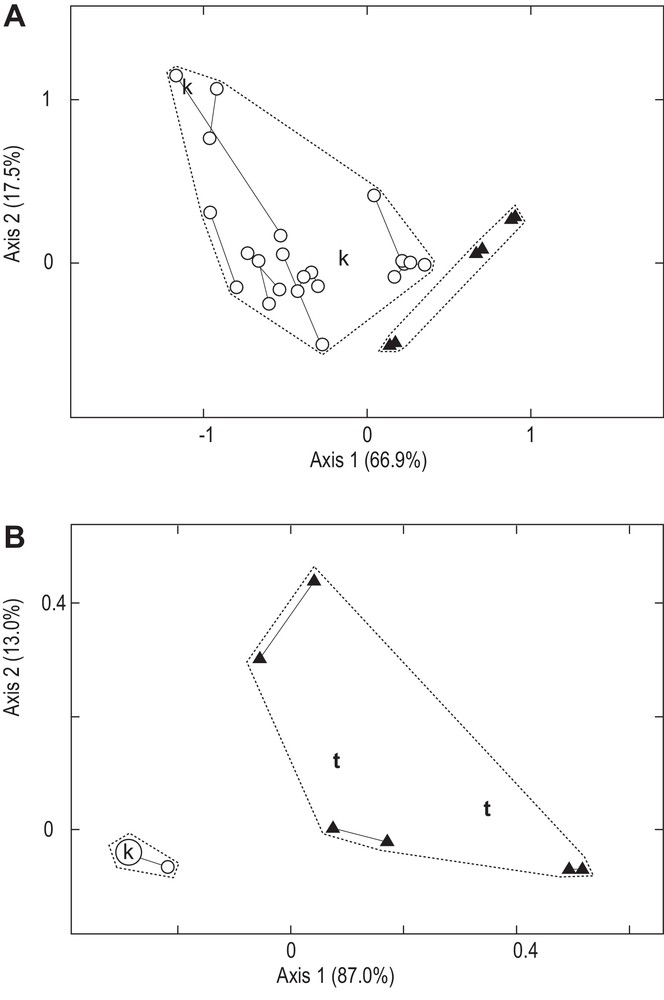
Neotrygon kuhlii and Neotrygon trigonoides. Correspondence analysis (CA: [28]) of the matrix of individuals characterized by the size-frequencies of ocellated spots on the dorsal side of a half-disk (Table 2). CA was run using the FactoMineR package [29] under R [30]; percentages for each axis are their inertias [28]. Open circles (○): N. kuhlii; closed triangles (): New Caledonian maskrays, N. trigonoides; k: N. kuhlii specimen depicted in Plate 51 of [9]; t: holotype of N. trigonoides. Thin segments link data points representing the two half-disks of an individual. Dotted lines delineate groups of individuals grouped by hierarchical clustering analysis [29]. A. Individuals characterized by six variables; N. trigonoides holotype excluded. B. All 40 individuals characterized by three variables (number of black speckles, number of black dots and presence/absence of brown scapular blotch), to allow the placement of the holotype of N. trigonoides in the absence of ocellated spot data.
4 Discussion
Contemporary species concepts are diverse but all relate to the idea that species more or less can be defined as segments of lineages of the global genealogical network pertaining to the metapopulation level of organization [31]. Properties of species may include reproductive isolation from other species, the possession of fixed character state differences, and monophyly [31–34].
Neotrygon trigonoides was originally defined as a distinct species from its unique spotting patterns [16]. The reason for declaring N. trigonoides a synonym of the distinctly colour-patterned N. kuhlii originates from the similarity of the N. trigonoides holotype, from New Caledonia, with blue-spotted maskray specimens from eastern Australia [17] which for some unclear reason had been erroneously identified as N. kuhlii. As shown in this paper, N. trigonoides should be rehabilitated as a valid species and blue-spotted maskrays from eastern Australia are of this species.
Haplotypes from blue-spotted maskray specimens sampled from the northern Great Barrier Reef and from New Caledonia (Coral-Sea) clustered into a well-defined and statistically supported clade external to N. kuhlii in the NJ tree of genetic distances. Character-evolution based ML phylogeny also found Coral-Sea haplotypes to cluster externally to those of N. kuhlii, although it failed to group them within a truly monophyletic haplogroup. We interpret this minor difference with the topology of the NJ tree as an indication that the phylogenetic information from the single and short COI fragment sequenced in this study might be insufficient. M. Puckridge et al. [15] have recently produced a phylogeny of Neotrygon spp. based on concatenated partial COI and 16S haplotypes, which confirms this hypothesis. All blue-spotted maskray mitochondrial haplotypes from the Coral-Sea (including new individuals from New Caledonia, from the northern Great Barrier Reef, from southeastern Queensland and from northeastern New South Wales) clustered as a well-defined clade in Puckridge et al.’s [15] phylogenetic reconstruction (their figure 2b). This was resolved as a clade sister to that formed by N. kuhlii mitochondria [15]. The N. kuhlii clade included a specimen from the Gulf of Carpentaria, which is geographically adjacent to the northern Barrier Reef where only N. trigonoides haplotypes have been sampled thus far ([8,15]; present results). This abrupt geographic discontinuity in mitochondrial genetic composition relative to the genetic continuity observed all across the Coral-Sea on the one side, and the Sahul shelf and even the whole Indian Ocean and Coral Triangle on the other side ([15], present results) is incompatible with the idea of a gradual genetic transition between N. trigonoides and N. kuhlii. Because no contemporary discontinuity of habitat suitable to N. kuhlii [10] is apparent across the Torres Strait region, this points to reproductive isolation between the two species.
Although reproductive isolation was here inferred between N. trigonoides and N. kuhlii, no firm proof of this has been presented. Relying on the single mitochondrial phylogeny may appear insufficient for assessing systematic relationships. However, nuclear (RAG-1) haplotypes from either side of the Torres Strait are also different [15]. Another line of evidence for two separate species is the large nucleotide distance between N. trigonoides and N. kuhlii, which was comparable to the distance between N. leylandi and N. picta. Last, the colour pattern of N. trigonoides distinguishes it from Müller and Henle's [9] N. kuhlii making it diagnosable on the basis of trivial external examination. Thus, N. trigonoides should be resurrected as a valid species because of its diagnosability when using both colour patterns and genetic characters, because of the monophyly of its mitochondrial DNA haplotypes, and because of the geographic distribution of haplotypes which highlights the genetic gap between N. kuhlii from the Gulf of Carpentaria (west of Torres Strait) and N. trigonoides immediately east of it. All the foregoing constitutes properties expected from separately evolving metapopulation lineages, i.e., distinct species [31].
The COI nucleotide sequences of N. kuhlii examined in this paper were sampled from 34 specimens from 18 locations spanning almost the full longitudinal range of the species, from the eastern coast of Africa to the West Pacific. Several well-supported sub-clades were visible within the N. kuhlii clade as observed previously [3,7,15]. The occurrence of deeply rooted clades within N. kuhlii has led authors to suggest that it consists of several cryptic species [3,15]. In the present study, as in other COI-based studies [7,8], we find no compelling, definitive evidence favouring this interpretation (Fig. 3). Likewise, spotting patterns as measured in our study did not allow a clear partition of N. kuhlii into distinct groups. However, more detailed nuclear-based investigations are needed, to test whether different mitochondrial clades within N. kuhlii may belong to reproductively isolated populations. The RAG-1 marker has proven to be too conservative to this effect [15]. Size-polymorphic introns [14] might be more appropriate. The few intron data available thus far [14] are compatible with a discontinuous distribution of genetically differentiated, possibly isolated populations. However, denser geographic sampling is necessary to further investigate potential geographic boundaries or transition zones.
5 Resurrection of Neotrygon trigonoides
Family Dasyatidae Jordan, 1888 [35]; Dasyatidae belong to the superfamily Myliobatoidei recently redefined on the basis of mitochondrial genomes and nuclear genes [1]. Genus Neotrygon Castelnau, 1873 [16]. N. trigonoides Castelnau, 1873, resurrected species.
5.1 Previous references
Raya trigonoides[16]; Dasyatis kuhlii (non Müller & Henle, 1841) [36,37]; N. kuhlii (non Müller & Henle, 1841) [8,17,38]; N. trigonoides [5,16].
5.2 Material examined
The N. kuhlii and N. trigonoides material examined for nucleotide sequence variation at the COI locus is listed in Table 1. Specimens of N. kuhlii and N. trigonoides examined for spotting patterns are listed in Table 1 and their photographs are presented in Supplementary material, Figs. S1 and S2, respectively.
Three vouchers of N. trigonoides that were characterized by their nucleotide sequences at the COI locus were placed in ichthyological collections: CSIRO uncatalogued, female, 225 mm DW, from Saint-Vincent Bay, New Caledonia – this specimen was captured on 16 August 2008 by P. Morlet and deposited in the Australian National Fish Collection (CSIRO, Hobart) by P.B. on 14 August 2009 (Supplementary material, Fig. S1); MNHN 2009-0823, female, 350 mm DW, New Caledonia (Fig. 1D); IRDN 20090816, male, 302 mm DW, from Saint-Vincent Bay, New Caledonia, captured on 16 August 2009 by P. Morlet and deposited in the fish collections of the IRD centre in Nouméa, New Caledonia.
5.3 Redescription
Castelnau [16] has provided a qualitative description of the colour patterns of N. trigonoides: “Entirely of a light brown lilac colour, with a few faint white oscillated [sic] spots on the disk, and a larger number of smaller black ones dispersed in a most irregular way; posterior part of the tail annulated, black and orange; lower side of the body entirely of a light cream colour”. The holotype of N. trigonoides (NMV 51684; Supplementary material, Fig. S1), a juvenile male, 182 mm DW, from New Caledonia, seems to have lost its ocellated spots in preservative. Live or freshly captured N. trigonoides are characterized by the presence of dark spots larger than > 1% DW and by a symmetrical pair of brown blotches in the scapular region (Supplementary material, Fig. S2).
Although spotting patterns are useful as a character to distinguish between N. trigonoides and N. kuhlii (Fig. 4), the mitochondrial sequence has proven to be an excellent diagnostic character (Fig. 3) and because of this fact, we consider it to be much more adequate than any of the morphological characters employed thus far, including spotting patterns (present work), to base our description on.
The present redescription of N. trigonoides is based on the nucleotide sequence of a 655-base pair (bp) fragment of the COI gene (hereafter abbreviated as ‘partial COI gene’) homologous to the portion of the mitochondrial genome of N. kuhlii comprised between nucleotide sites 2278 and 2932 (GenBank JN184065; [1]). The partial COI gene of specimen MNHN 2009-0823 has the following sequence (accession no. JX263420 in GenBank): 5′- C C T T T A C T T A G T C T T T G G T G C A T G A G C A G G G A T A G T A G G C A C T G G C C T T A G T T T A C T T A T C C G A A C A G A A C T A A G C C A A C C A G G C G C T T T A C T G G G T G A T G A T C A A A T T T A T A A T G T A A T C G T C A C T G C C C A C G C C T T C G T A A T A A T C T T C T T T A T G G T A A T G C C A A T T A T A A T C G G T G G G T T T G G T A A C T G A C T A G T A C C C C T G A T G A T T G G A G C T C C G G A C A T A G C C T T T C C A C G A A T A A A C A A C A T A A G T T T T T G A C T T C T A C C T C C C T C C T T C C T A C T C C T G C T A G C C T C A G C A G G A G T A G A A G C T G G A G C T G G A A C A G G T T G A A C A G T T T A T C C C C C A T T A G C T G G T A A T C T A G C A C A T G C C G G A G C T T C T G T A G A C C T T A C A A T C T T C T C T C T T C A C C T A G C A G G T G T C T C C T C T A T T C T G G C A T C C A T C A A C T T T A T C A C A A C A A T T A T T A A T A T A A A A C C A C C T G C A A T C T C C C A G T A T C A A A C C C C A T T A T T C G T C T G A T C C A T T C T T G T T A C A A C T G T A C T T C T C C T G C T A T C C C T A C C A G T C C T A G C A G C T G G C A T T A C C A T A C T C C T T A C A G A C C G A A A T C T T A A C A C A A C T T T C T T T G A C C C A G C T G G A G G A G G A G A T C C C A T T C T T T A C C A A C A T C T C T T C -3′. This sequence has accession No. KC295416 in GenBank.
In addition, the nucleotide sequence of the cytochrome b gene (1132-bp) of N. trigonoides MNHN 2009-0823 homologous to the portion of the mitochondrial genome of N. kuhlii comprised between nucleotide sites 10859 and 11992 (GenBank JN184065; [1]) is the following: 5′- A A C A T C C G T A A A A C A C A T C C C C T A T T C A A A A T T A T C A A C A A C T C A C T A A T T G A T C T A C C A G C T C C A A C C A A T A T T T C C A C C T G A T G A A A T T T T G G T T C C C T A C T A G G C C T T T G C C T A A T T A T C C A A A T C C T T A C A G G C C T A T T C C T A G C T A T A C A C T A C A C C G C A G A C A T C T C A T C A G C A T T C T C C T C A G T T G C C C A T A T C T G C C G A G A C G T T A A C T A C G G T T G A C T A A T C C G C A A T A T T C A C G C T A A C G G C G C C T C A A T A T T C T T C A T C T G T G T T T A T C T C C A T A T T G C T C G A G G A C T T T A C T A T G G C T C C T A C C T C A A T A A A G A A A C C T G A A A T A T C G G A G T A G T T A T C C T A G T G T T A C T A A T A G C C A C C G C A T T C G T A G G C T A T G T T C T C C C A T G A G G A C A A A T A T C A T T C T G A G G G G C A A C C G T T A T C A C C A A C T T A C T A T C A G C C C T C C C C T A T A T T G G A G A C A T G T T A G T T C A A T G A A T C T G A G G G G G C T T C T C A A T T G A C A A T G C A A C A T T A A C T C G A T T T T T C A C A T T T C A T T T T C T A T T T C C C T T T G T A A T T G C A G C T C T T A C T A T A A T T C A C C T T C T C T T C C T T C A T G A A A C A G G T T C T A A C A A C C C A A C C G G A C T C T C A T C T A A C A T A G A C A A A G T C C C G T T T C A T C C T T A T T A T A C A T A T A A A G A T C T A G T A G G C T T C T T C A T C C T T C T A A T A C T A C T A A C T C T A C T T G C C C T A T T T A C A C C A A A C C T C C T A G G G G A T A C A G A A A A C T T T A T T C C A G C C A A C C C C C T C G T C A C A C C T C C C C A T A T T A A A C C A G A G T G A T A C T T C T T A T T T G C C T A C G C T A T T C T A C G C T C T A T C C C C A A T A A A C T A G G A G G A G T C C T A G C C C T C G C C T T C T C A A T C T T T A T C C T G C T A C T A A T C C C C A T T C T T C A C A C C T C T A A A C A A C G A A G C C T T A C C T T C C G T C C A A T T A C A C A A C T C C T G T T C T G A C T C T T A G T G G C A A A C A C A A T C A T C C T A A C A T G A A T C G G C G G C C A A C C C G T A G A A C A G C C A T T C A T C A T T A T T G G C C A A A T C G C C T C A A T C A C C T A C T T C T C C T T C T T C C T C A T C C T A T T C C C A A T C G C T G G A T G A T G A G A A A A C A A A A T G T T A A A C C T T A -3′. This sequence has accession no. KC493691 in GenBank.
5.4 Vernacular names
N. trigonoides is apparently endemic to the Coral-Sea. As the holotype of the species is from New Caledonia, and as to our knowledge there is no Neotrygon species other than N. trigonoides occurring in New Caledonia, we propose as the English vernacular name: New Caledonian maskray. We propose as the French vernacular name: raie pastenague à points noirs et bleus, as it is usually called in New Caledonia [39], even though gris (grey) might be a more accurate epithet than bleus (blue) to designate the colour of the ocellated spots.
5.5 Comparison with closely related species
Species of the genus Neotrygon (N. kuhlii, N. leylandi, N. ningalooensis, N. picta and N. trigonoides) show no diagnostic difference in the partial COI amino-acid sequence as translated from the 519-bp fragment analyzed in common in all five species (present study). Among species in the genus, N. trigonoides is most closely related to N. kuhlii (Fig. 3). N. trigonoides can be separated from N. kuhlii by the nucleotide synapormorphies highlighted on Fig. 5.
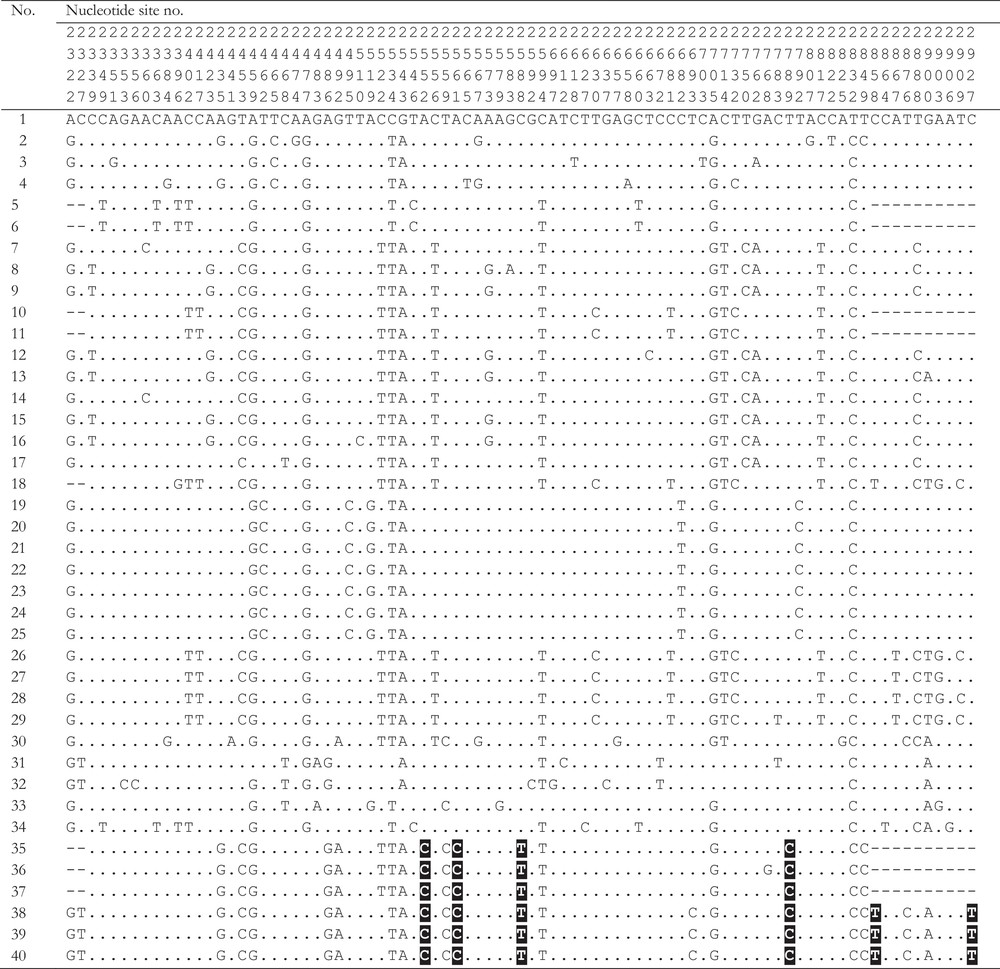
Variable nucleotide sites in COI gene sequences of Neotrygon kuhlii and Neotrygon trigonoides. Numbers allocated to specimens as in Table 1 and Fig. 2. Nucleotide sites numbered according to [1] (GenBank JN184065). Highlighted: synapomorphies characterizing the New Caledonian maskray, N. trigonoides.
5.6 Geographic distribution
DNA-barcoded N. trigonoides specimens were collected in Lizard Island [8], off southeastern Queensland and northeastern New South Wales [15] and in New Caledonia ([15], present study). Therefore, based on the material genetically identified to species thus far, N. trigonoides exclusively occurs in the Coral-Sea. Blue-spotted maskrays from Moreton Bay (southeastern Queensland) in the southwestern Coral-Sea also possess dark spots and the symmetrical pair of scapular blotches on either side of the chord characteristic of N. trigonoides (although the latter are fainter than in specimens from New Caledonia); however, in contrast with New Caledonian specimens, their ocellated spots are pale-blue and are also larger in size relative to DW (from a series of photographs by S. Theiss; pers. comm.). Based on spotting patterns from specimens photographed underwater by F.R. McConnaughey (www.gettyimages.com), N. trigonoides is also present in Vanuatu at the eastern boundary of the Coral-Sea [40].
Disclosure of interest
The authors declare that they have no conflicts of interest concerning this article.
Acknowledgements
We are grateful to Susan Theiss (University of Queensland, Brisbane) for sharing excellent pictures of blue-spotted maskrays from Moreton Bay, and to Hans Ho (NMMB, Pingtung) for sub-sampling tissue from a specimen from Taiwan. Gavin Naylor and Zhong-Duo Wang kindly provided sampling details for some of the Neotrygon spp. material referred to in the present study. We are grateful to Patrick Morlet, fisherman in Bouraké, New Caledonia, for fishing and preserving N. trigonoides specimens for us. We thank Catherine Picq (IRD, Montpellier) for her help in getting a copy of the original description by Castelnau; Renny Hadiaty (MZB, Cibinong) and Patrice Pruvost (MNHN, Paris) for allocating collection numbers to our specimens; and Dianne Bray (Museum Victoria, Melbourne) for details on the holotype of N. trigonoides. The opus of Müller and Henle was consulted online from the Biodiversity Heritage Library Internet site (http://www.biodiversitylibrary.org). Conceived and designed the experiments: PB. Contributed reagents/materials/analysis tools: PB, WJC, ISA, JDD, MGM, KNS. Did the experiments: PB, ISA, KNS. Analyzed the data: PB, WJC. Wrote the paper: PB. Funded by AIRD, IRD, ISE-M, LIPI-P2O, and National Taiwan University-Institute of Fisheries Science; the funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.


