1 Introduction
Wood is a renewable natural resource, providing timber (e.g. for house building, furniture, packaging), fibres (for pulp, paper, plywood) and energy (firewood). Recently, considering the widespread depletion of fossil energy resources, wood has become a potential source of lignocellulose-based biofuels. A product of high economical value, wood represented a major advance in plant evolution during the Cretaceous and allows trees to reach considerable heights and lifespans. The major functions of wood are to:
- (1) conduct water from roots to the crown;
- (2) support an ever-increasing mass of the growing tree, whilst adjusting to various environmental cues (wind, snow, slope, light);
- (3) contribute to tree growth over more than one year by storing temporary reserves.
In angiosperm trees, different cell types fulfil these three functions. Vessels and fibres are involved, respectively, in water conduction and mechanical support, while parenchyma cells, organized in rays, are involved in the radial transfer of assimilates between phloem and xylem, their temporary storage as starch or lipids, and their remobilization at the new season. Angiosperm wood is therefore heterogeneous. In the homogeneous wood of gymnosperms and some primitive angiosperms, the tracheids ensure both water conduction and tree support. In this review, we will focus on wood formation in angiosperms.
Wood formation is a complex developmental process, involving five major developmental steps, including
- (1) cell division from a secondary meristem called the vascular cambium;
- (2) cell expansion (cell elongation and radial enlargement);
- (3) secondary cell wall deposition;
- (4) programmed cell death;
- (5) heartwood formation.
It has been extensively reviewed in terms of developmental biology [1–3]. In this review, we will first describe these different developmental steps, and then summarize the recent advances in deciphering molecular mechanisms underlying wood formation, thanks to the development of genomics on woody species. Two major wood features will be emphasized: the lignification process and the control of cellulose microfibril angle in the cell wall.
2 Wood formation, a complex developmental process
2.1 Wood is produced by the vascular cambium
Wood, also called secondary xylem, is produced seasonally at the periphery of the trunk by the vascular cambium. Derived from the procambium, this meristem is responsible for shoot and root secondary growth [4]. It is of major importance in the perennial life of trees as it regularly produces functional xylem and phloem cells, towards the inner and outer parts of the trunk, respectively. The cambium is made of initials, which divide to produce phloem and xylem mother cells [4]. When it is reactivated in the spring, in many species, the cambium starts to produce first phloem cells, sometimes several weeks before any xylem cells are produced. However, at the end of the growing season, the number of xylem cells formed exceeds the number of phloem cells [5]. Wood is therefore produced in a larger quantity than secondary phloem in the stem. In contrast to primary meristems, the cambium is made of two types of initials, different in size and shape. Elongated fusiform initials give rise, by periclinal divisions in the tangential plane, to the axial cell system in wood (water-conducting vessel elements, vessel-associated cells, fibres and axial parenchyma cells) while radial isodiametric initials produce the radial cell system, consisting of the parenchyma ray cells [2,6,7] (Fig. 1B). The identity of the cambial cells seems to be linked to positional signals, more than to the identity of the cell lineage, as fusiform and radial initials can inter-convert [4]. On a stem cross-section, the cambial zone is characterized by a few layers of narrow and thin-walled cells (Fig. 1B). Anticlinal divisions of fusiform initials ensure the harmonious increase in the cambium circumference.
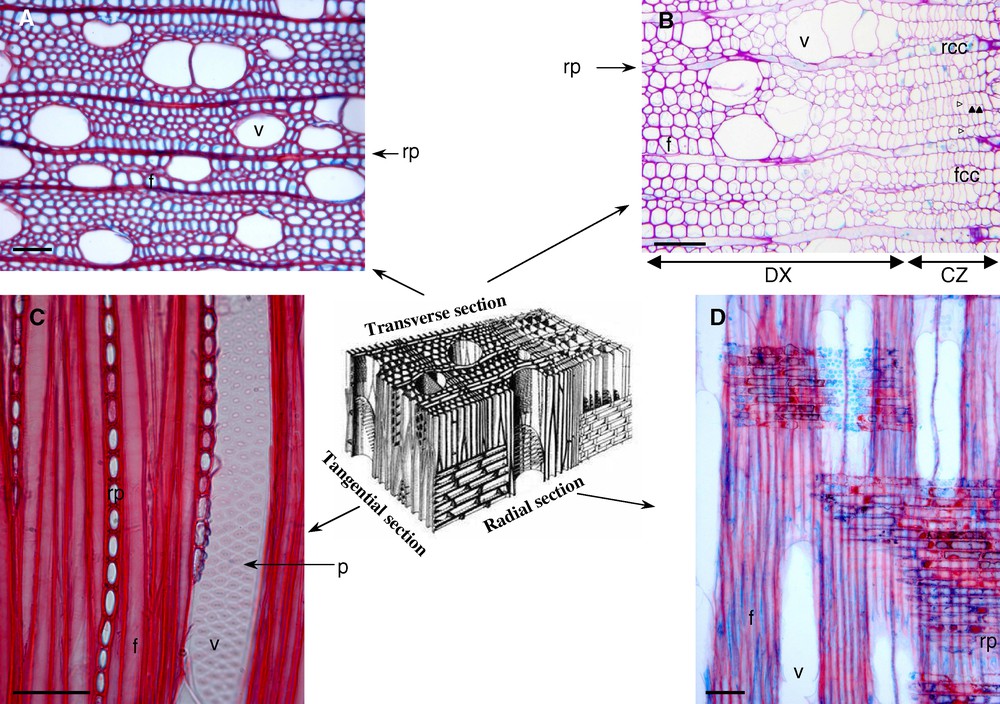
Wood results from the three-dimensional association of different cell types, as represented on the central scheme (adapted from [91]) Transverse (A, B), tangential (C) and radial (D) sections from poplar (Populus tremula x P. alba, INRA clone 717-1B4), stained with safranine/Astra blue. CZ: cambial zone; DX: differentiating xylem; f: fibre; fcc: fusiform cambial cell; p: pit; rcc: radial cambial cell; rp: ray parenchyma; v: vessel. Empty arrowhead indicates a newly deposited tangential wall (periclinal division) in the cambial zone. Full arrowhead indicates a newly deposited radial wall (anticlinal division) in the cambial zone. The scale bar corresponds to 50 μm.
2.2 Wood cells undergo a profound differentiation ending by programmed cell death
At the beginning of their differentiation, these young xylem cells lengthen without increasing width. At this stage, they have only a thin primary cell wall. Then, they increase in diameter, set up their secondary wall and acquire the distinctive characteristics of their cell type. For example, the vessels are easily distinguishable from fibres owing to their larger diameter (Fig. 1). The axial expansion of the vessel elements is however limited, in comparison to fibres (Fig. 1C). The walls connecting vessel elements are perforated, leading to the formation of a tube, adapted for water conduction over long distances [8]. The vessel will deposit a secondary cell wall thickening, but showing pits without secondary cell wall on the longitudinal sides, allowing communication, water and nutrient transport with the adjacent vessel element or parenchyma cell (Fig. 1C and D). In contrast, the fibres elongate dramatically in the axial direction, while keeping a very moderate radial expansion. They can reach 0.6 mm length in poplar, 0.9 mm in black locust, 1 mm in oak and chestnut, 1.5 mm in beech [9,10]. Beside symplastic growth, fibres have an intrusive growth, with local cell wall biosynthesis in the fibre tips. Recently, transgenic hybrid aspen trees with modified pectin methyl esterase activity supported the hypothesis that homogalacturonan methylesterification is involved in cell wall plasticity [11]. Actin filaments, aggregated to form cables, are thought to be involved in the transport of secretion vesicles towards the fibre tips in these highly vacuolated cells; they are distinctly oriented along the fibre axis, in poplar and chestnut [6]. This process of intrusive growth is very similar to the growth of pollen tubes or root hair.
When they reach their final size, the fibres deposit a very thick secondary cell wall. The secondary cell wall of both fibres and vessels will be lignified. The lignification of the walls starts from the cell corners, and progress from the middle lamella to the primary and secondary walls. Lignin deposition starts before the entire layers of secondary cell wall are deposited [12]. Lignins associate gradually with hemicelluloses present in the cell wall, eliminate water and form a hydrophobic environment. The cells become both rigid and impermeable.
Finally, the cells undergo programmed cell death. The vacuole membrane dissociates and releases specific hydrolases (Cys and Ser proteases, nucleases and RNases) [13–16]. In addition, cytosolic pH dramatically decreases, activating cytosolic hydrolases. Nucleus, plastids, mitochondria disappear, and the cells are emptied of their cytoplasm. Note that the parenchyma cells, despite a lignified secondary cell wall, can remain alive and functional for several years, showing that programmed cell death is not directly connected to lignification. The parenchyma ray cells are involved in temporary reserve storage during the winter, as well as in the inter-connection between secondary phloem and xylem.
However, when these parenchyma ray cells become dysfunctional, they die, liberate phenolics, and contribute to the formation of heartwood, the most internal part in the trunk, which often appears dark-coloured, in comparison to sapwood. Sapwood, the most external part in the trunk under the bark, is the living part of the trunk where water conduction is effective. In between, a transition zone can be detected, where heartwood generation and extension occurs, from the centre of the trunk toward its periphery. The released phenolics, synthesized in parenchyma cells, diffuse in the neighbouring cell walls and lumens, giving heartwood its specific colour and also, potentially its increased long-term resistance to pathogen attack (durability). For many hardwood species, heartwood formation is preceded by vessel plugging: the vessels become occluded by tyloses, balloon-like outgrowths developing from parenchyma cells via the pits, impeding water transport [5]. Heartwood characteristics vary greatly according to tree species and are responsible for the main chemical and aesthetic properties of woods that will determine their final uses. For example, species like walnut and black locust produce heartwood that is much more coloured and durable than their sapwood. Poplar has weakly differentiated heartwood also called false heartwood that is only lightly coloured, and not durable.
2.3 Wood is mostly made of secondary cell walls of dead fibres
The secondary walls are made of several layers, called S1, S2, S3, each composed of a network of long bundles of cellulose microfibrils, oriented at a fixed angle, and cemented in an amorphous matrix of hemicelluloses – xylans mainly –, and lignins [17]. In smaller quantities, proteins and pectins are also present. The S1, S2 and S3 layers differ in their thickness, chemical composition and cellulose microfibril angle (MFA, measured from the fibre longitudinal axis) (Fig. 2A and B). In particular, the alternation of high and low angles in the different layers of the secondary cell wall increases its rigidity and mechanical strength. From a mechanical point of view, the cell wall can indeed be regarded as a composite material, like reinforced concrete [18]. Models derived from the mechanics of composite materials show the importance of MFA and cellulose crystallinity in explaining the macroscopic mechanical properties of wood [19]. MFA in the S2 layer, which is the thickest one (Fig. 2 A and B), is a parameter widely used in wood technology. As MFA increases – the cellulose microfibrils diverge more from the vertical – stiffness and tensile strength decrease [20]. An inverse relationship between MFA in the S2 layer and fibre length is generally observed for a given wood sample between growth rings: the closer the MFA is to 0°, the longer the fibre. However, fibres of the same length from different species can have widely different MFA [21].
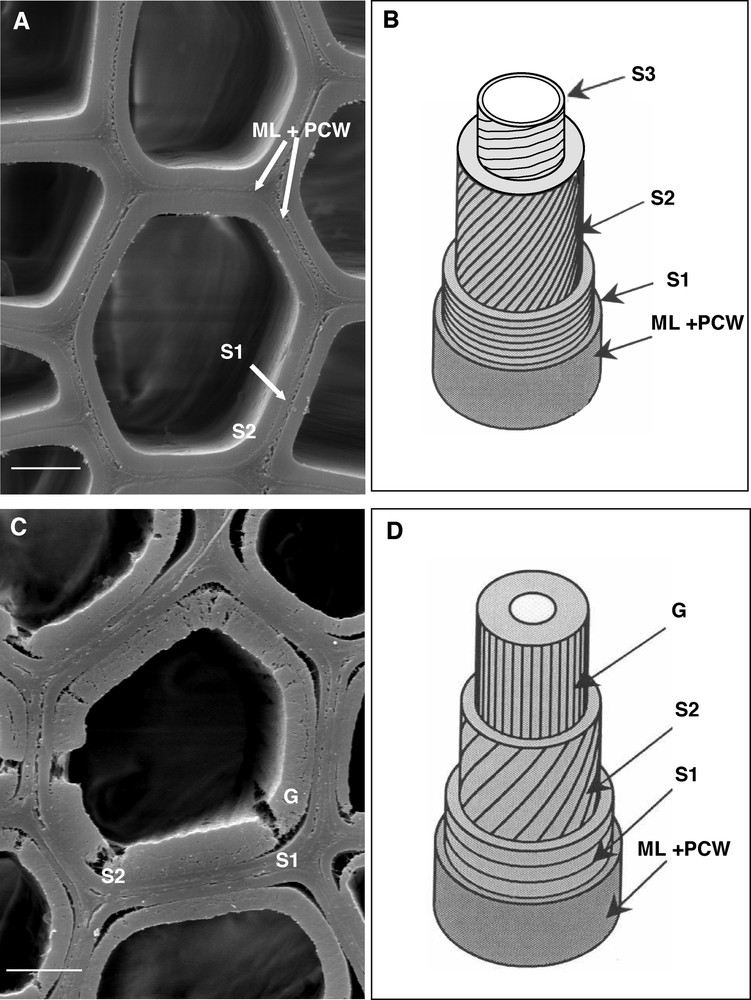

The secondary cell wall is composed of several identified layers. Scanning electron microscopy and interpretation scheme of the different cell wall layers of normal (A, B) and tension wood (C, D) fibres of poplar. Note that the lines on the scheme indicate the preferred orientation of cellulose microfibrils in the different layers. ML: middle lamella; PCW: primary cell wall; S1, S2, S3: layers of the secondary cell wall; G: gelatinous layer in tension wood fibers. The scale bar corresponds to 5 μm.
From a biochemical point of view, wood is composed of about 50 % cellulose, 25 % lignins and 25 % hemicelluloses, with traces of pectins and proteins, impregnated with phenolics. These values can vary a lot between tree species and between individuals in the same tree species (see the next section). Cellulose is for pulp or bioethanol production the valuable biochemical compound. To have access to cellulose microfibrils embedded in the cell wall's intricate structure, delignification treatment is often necessary. On the other hand, lignins and phenolics are important biochemical compounds for wood calorific properties.
2.4 Wood is very variable
Wood is a complex tissue made of the three-dimensional assembly of different cell types, characteristic of a given species (Fig. 3). While fibres constitute the most important part of wood, the proportion of parenchyma ray cells can vary a lot between tree species (5% for Tilia, 9% for Populus, up to 30% for Quercus [9]). The size and the number of vessels are also species-dependant. However, this typical wood pattern can be considerably modified under different factors. Within an annual ring, early wood produced in the spring when temperature and photoperiod are favourable for vegetative growth, differs anatomically from latewood, when cambium is less active (Fig. 3A and B). Late wood is denser due to smaller fibres with thicker cell walls and smaller lumen. There are also differences in the properties of juvenile wood, produced both in young trees (up to the first 5–20 years of its life) and in the youngest apical parts of older trees, and the properties of mature wood produced by old trees. Juvenile wood is characterized by a lower density and a higher MFA: the young tree can bend, for example under the wind, without breaking. In mature wood, the MFA is lower: the wood is denser, leading to a mechanically more resistant material, better suited for an utilisation in house building, for example.
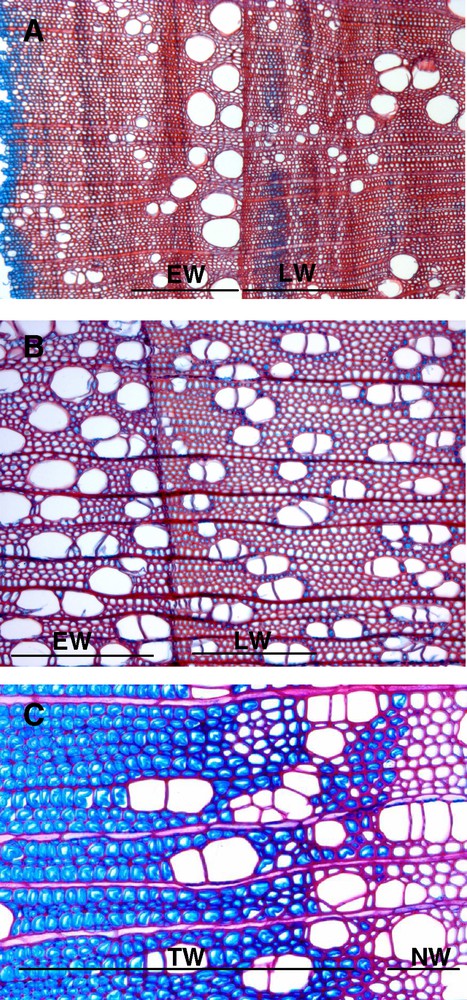
Wood is a very variable tissue. Transverse wood sections of chestnut (A) and poplar (B, C), stained with safranine/Astra blue, showing differences in vessel size, vessel number, fibre size, fibre secondary cell wall between earlywood (EW), latewood (LW), normal wood (NW) and tension wood (TW).
In response to certain environmental cues (wind, slope, light), the tree can develop a particular wood, called reaction wood, with specific mechanical properties, in order to reorient its axes, trunks and branches, and to maintain its general equilibrium [22]. In angiosperms, this wood, produced at the upper side of inclined axes, is named tension wood. At the macroscopic level, a high eccentricity is often observed in the transverse section of stem, suggesting that cell divisions are activated on tension wood side. At the microscopic level, tension wood has fewer vessels of smaller size than normal wood [23]. This wood is characterized remarkably by the differentiation of peculiar longer fibres, called G fibres, with a thick inner gelatinous layer, which can fill almost the whole fibre lumen (Fig. 2C, 2D, 3C). This G layer consists primarily of pure cellulose, organized in aggregates or macrofibrils of 30 to 40 nm diameter [24]; it also contains xyloglucans [25] and is almost free of lignins [26–28]. Cellulose microfibrils, much more crystalline than in other layers of the secondary wall are oriented in the direction of the axis of the fibre, with a MFA close to 0° [29] (Fig. 2C and D). Only 50% broadleaved species differentiate true G fibres in tension wood [30]. But in all cases, the tension wood fibres are characterized by a high level of crystalline cellulose with low MFA. These two features are important to explain the peculiar tension properties of this wood.
2.5 Control of wood formation
Cambial activity and xylem differentiation are controlled, at least partly, by different hormones [2]. An auxin concentration gradient was observed on poplar wood cross-sections, peaking in the cambial zone [31]. This has led to the hypothesis that auxin is required for cell division, and that its concentration level constitutes a positional signalling for cambial initial division and xylem cell differentiation. To support this hypothesis, it was shown that apically applied exogenous auxin could restore cambium activity in decapitated shoots [32,33]. More recently, using transgenic poplars down-regulated for auxin signaling, Nilsson et al. [34] demonstrated that auxin is required for cell proliferation and cell differentiation during cambial development. Besides auxin, several other hormones, including cytokinins [35], gibberellins [33], and ethylene [36], stimulate cambium cell division when applied exogenously, so they are potentially involved in the control of cambial activity in planta. Recently, transgenic poplar trees expressing a cytokinin catabolic gene in the cambial zone exhibited an effectively reduced level of cytokinins, and a reduced radial growth [37]. Using also a transgenic-based approach, Love et al. [38] showed that endogenous ethylene produced in leaning stems stimulated cambium cell division and could explain poplar stem eccentricity associated with tension wood formation. The hormonal control of wood formation seems to be rather complex and most probably involves cross talk between the different hormones, in response to developmental events and/or environmental cues.
The different types of wood described above can coexist within a single tree, showing that wood properties are highly variable. This plasticity of wood properties partly results from changes in the expression of specific genes/proteins during wood formation. Although it is a complex developmental process, it is possible to take advantage of its highly organised structure to develop a relevant genomics approach, in order to decipher the molecular mechanisms underlying each developmental step.
3 Recent advances in the molecular biology of wood formation
3.1 Development of genomics for wood biology
The first genomic programs concerning wood biology started more than 10 years ago, with the sequencing of the first 5,692 Expressed Sequence Tags (EST) of wood-forming tissues of Populus tremula x P. tremuloides and P. trichocarpa [39]. Today, more than 300,000 EST are available from 18 different poplar species, with almost 43,000 from wood tissues (source: NCBI dbEST, 09/09/23). As wood is spatially organized, it is possible to sample, extract RNA, prepare cDNA libraries and/or perform transcriptomics analyses from specific tissues. This has been done successfully, for example, for the cambial zone, developing xylem and mature xylem of poplar tension and opposite wood [40], for poplar fusiform and radial cambial initials [41], for bark/cambial zone, sapwood and transition zone from Robinia pseudoacacia [42]. EST sequencing is then powerful to discover genes involved in specific developmental events. For example, EST abundance between cambial zone and xylem specific cDNA libraries were compared, and this showed that there was a dramatic change in gene expression in poplar. In the cambial zone were mainly found EST involved in the cell cycle, protein synthesis and fate, while in xylem, most abundant EST clusters corresponded to genes involved in cellulose synthesis (sucrose synthase, cellulose synthase), methylation of lignin precursors (S-adenosylmethionine synthetase, homocysteine methyltransferase, caffeoyl-CoA O-methyltransferase –CCoAOMT-, caffeic acid/5-hydroxyconiferaldehyde O-methyltransferase –COMT-), cytoskeleton organization (α tubulins, actin-depolymerizing factor) [40]. This EST sequencing approach led to the identification of 10 new tension-wood specific genes, corresponding to fasciclin-like arabinogalactan proteins (FLA, [43]). These genes have only one Arabidopsis homolog, AtFLA12, indicating that this gene family has expanded a lot in poplar. FLA proteins are very abundant in the G layer of tension wood fibres and were specifically detected at the interface with plasmalemma [43]. These proteins have a fasciclin-like domain, which is putatively involved in protein-protein interactions. Although the role of these highly glycosylated proteins is not fully understood, they are most probably involved in the structural organization of the G layer. Mellerowicz et al. [44] suggested that FLA may create the required hydrophilic environment in the G layer, necessary for Xyloglucan Endo Transglycosylase (XET) to function. According to these authors, this enzyme is involved in the cohesion of G layer to the surrounding secondary cell wall.
The development of molecular biology tools to study heartwood formation is quite recent due to technical limitations – low number of living cells and high phenolic content within the oldest part of the trunk, especially in the sapwood/heartwood transition zone [45,46]. However, EST sequencing programs have been developed or are currently under development for black locust [41] and walnut. They first highlighted the up-regulation of flavonoid biosynthesis genes in the transition zone, which was in agreement with previous studies. For example, in black walnut (Juglans nigra), strong flavanol accumulations are correlated with high transcript levels of chalcone synthase and flavanone 3-hydroxylase genes in transition zone during the heartwood expansion period [47]. In this species, the oxidation and polymerization of juglone (a very reactive naphtoquinone specific of Juglandaceae) may explain in great part the heartwood colouring processes. However, its production and biosynthesis pathway remain quite complex and β-glucosidases may represent a key factor of juglone synthesis in the sapwood/heartwood transition zone.
Large transcriptomic analyses were developed on woody species, setting the basis for a comprehensive analysis of gene networks [48–50]. These large-scale studies can help to identify regulators in the process of wood formation. For example, transcriptomics underlined that ethylene and auxin-responsive genes were up-regulated, during poplar tension wood formation [51], suggesting that these hormones are involved in this developmental process. In addition, it identified transcription factors whose transcripts were up-regulated in tension wood: they are therefore potential key-regulators of tension wood formation.
For the last 15 years, poplar has become a model species for plant biology [52], first of all because it grows rapidly, can be propagated easily by cuttings, and can be genetically transformed using Agrobacterium tumefaciens, at least for given poplar clones [53]. In addition, because of the relatively small size of its genome (485 Mb versus 120 Mb for Arabidopsis), Populus trichocarpa was selected as the third plant species to be sequenced, after Arabidopsis thaliana and Oryza sativa. This sequencing was performed by the International Poplar Genome Consortium led by the U.S. Department of Energy (http://genome.jgi-psf.org/Poptr1_1.home.html) [54]. This was a large breakthrough for the poplar research community as it gave easy access to multigene families, to promoter regions, and facilitated comparison between poplar and Arabidopsis gene orthologs. Under chosen growing conditions, Arabidopsis can undergo secondary growth, and can therefore be an interesting genetic model in support to studies on trees, as mutants are easily available [55–57].
3.2 Lignification
Lignin is the second major polymer after cellulose in the plant secondary cell wall and its biosynthesis has been extensively studied, especially in tree species. However, it is considered as an undesirable compound for several uses of wood, like pulp or bio-fuel production. Lignin is synthesized from the oxidative coupling of p-hydroxycinnamyl alcohol monomers, called monolignols, differing in the number of methoxylation on their aromatic cycle, and other related compounds [58]. After lignin monomers are biosynthesized, they are translocated to the cell wall, where they are oxidized by peroxidases and/or laccases for polymerization. Angiosperm lignins mainly contain guaïacyl (G) and syringyl (S) units, deriving respectively, from coniferyl and sinapyl alcohols. These polymers, covalently bound to hemicelluloses, provide strength and rigidity to the cell wall, and hydrophobicity to the vascular system. In the past two decades, lignin biosynthesis has been better understood, thanks to the studies of mutant and/or transgenic plants altered for the expression of individual genes of the monolignol biosynthetic pathway. Many reviews have considered the details of the lignin biosynthetic pathway, including the impact of genetic engineering on it [58–61]. The general picture that emerges is that down-regulation of phenylalanine ammonia-lyase (PAL), cinnamate 4-hydroxylase (C4H), 4-coumarate: CoA ligase (4CL), p-hydroxycinnamoyl-CoA: quinate shikimate p-hydroxycinnamoyltransferase (HCT), p-coumarate 3-hydroxylase (C3H), CCoAOMT, cinnamoyl-CoA reductase (CCR), and, to a lesser extent, cinnamyl alcohol dehydrogenase (CAD), induced a decrease in total lignin content (for recent review, see [62]). Lignin composition (S/G ratio) can be also affected. For example, over-expression of ferulate 5-hydroxylase (F5H) can increase the S/G ratio in plants [63], while COMT down-regulation decreased the S/G ratio [64]. These different studies on transgenics also underlined that lignins can clearly incorporate other molecules than the classical three monolignols p-coumaryl, coniferyl, and sinapyl alcohols. COMT downregulation led to the incorporation of 5-hydroxyconiferyl alcohol into the lignin polymer; sinapaldehyde was incorporated in lignins when CAD was down-regulated [65–67]. These two molecules both resulted from incomplete monolignol biosynthesis. Lignins are therefore very variable polymers.
The availability of genomic resources (ESTs, genome sequence, microarrays) has widened the lignin research field. Raes et al. [68] identified the complete set of genes involved in monolignol biosynthesis pathway in Arabidopsis, using both the in silico approach and expression data. Most lignin biosynthesis gene families expanded rapidly and recently – after the divergence between monocots and dicots – [69]. This expansion correlates with substrate diversity. Current studies tend to take into account whole gene-family members in order to associate each member with specific processes thanks to co-expression analyses of genes. Lignin mutants or transgenics are now comprehensively analysed by transcriptome and metabolome profiling. Clearly, altering the expression of one gene in the monolignol biosynthetic pathway can have larger, and sometimes unexpected, consequences, on plant metabolism. For example, CCR-down-regulated transgenic poplars deposit less lignin, as anticipitated from previous research on the monolignol biosynthetic pathway in other species, but transcriptome and metabolome analyses also indicated a reduced xylan biosynthesis [70].
Transcript profiling of lignin transgenics is a way to identify new genes whose function is potentially closely associated with lignin biosynthesis, as well as regulatory transcription factors. Up to now, relatively few regulators, belonging to MYB and LIM family transcription factors, have been experimentally identified through both in vitro promoter binding experiments and transgenic approach in Eucalyptus [71–73]. In Arabidopsis, MYB58 and MYB63 have been demonstrated to be direct activators of the lignin biosynthetic pathway [74]. They are both downstream components in the SND1-mediated transcriptional network. SND1 (also called NST3/ANAC012) belongs to a group of closely related NAC domain transcription factors, together with NST1, NST2, VND6 and VND7. They have been shown to regulate a battery of downstream transcription factors, which in turn activate the biosynthetic genes for secondary wall deposition [75–78].
3.3 What controls MFA in the secondary cell wall?
MFA is controlled for a type of cell wall or S layer (Fig. 2); it is a critical parameter of the mechanical properties of wood. Very little is known about the control of MFA in trees, the most advanced studies arising from the model plant Arabidopsis (for recent review, see [79]). Cellulose is a linear chain of β 1-4 linked glucoses, synthesized by a complex multimeric transmembrane cellulose synthase (CesA), forming a 6-lobe structure, called the rosette, at the plasmalemma surface of plant cells. This rosette was clearly observed by electron microscopy [80]. It is believed that each rosette synthesizes simultaneously 36 cellulose molecules. The parallel chains of cellulose are likely to form many hydrogen bonds between them, leading to their crystallization as rigid insoluble rods, the cellulose microfibrils. The elongation of cellulose molecules can occur through the movement of CesA complexes inside the plasma membrane.
In various biological systems, it has been observed very frequently that cortical microtubules in the cytoplasm were oriented parallel to cellulose microfibrils in the cell wall [81]. These observations led to the hypothesis that microtubules were involved in the alignment of cellulose microfibrils, either actively, serving as molecular rails, or passively by forming channels limiting lateral movement of cellulose synthase rosettes. Paredez et al. [82] have clearly demonstrated by cell wall dynamic studies that in the primary cell wall of elongating Arabidopsis hypocotyl cells, the cellulose synthase complexes moved along trajectories defined by the cortical microtubules. However, if the microtubules were artificially disorganised, these complexes did not move randomly, suggesting that there is a part of physical self-organisation of the cell wall [83]. In vessels of primary xylem showing localized annular secondary wall thickenings, microtubules were also involved in the containment of CesA complexes in the plasma membrane areas underlying secondary wall rings [84]. These same authors also showed that actin cables, oriented along the axis of the vessels, were essential to cytoplasmic trafficking of organelles containing cellulose synthases to the sites of secondary wall synthesis. In addition, other molecular actors were identified thanks to the study of Arabidopsis fragile fibre mutants: a katanin-like protein encoded by FRA2 gene [85] and a kinesin-like protein encoded by FRA1 gene [86]. Both FRA1 and FRA2 mutations reduced stem growth and disorganised both cortical microtubule and cellulose microfibril networks. FRA2 codes for a katanin-like protein: in animal cells, katanin has been shown to possess a microtubule-severing activity. FRA1 encodes a kinesin-like protein. Kinesins are microtubule-binding motor proteins; it appears likely that FRA1 mediates the activity of microtubules in orienting cellulose microfibrils during differentiation of xylem cells in Arabidopsis [86].
In the case of secondary xylem fibres with a thick secondary wall, cortical microtubules may play a similar role in the control of MFA. Tension wood is a relevant biological model to study MFA control, as very strikingly, MFA is close to 0° in the thick cellulose-rich G layer (Fig. 2C and D). At the anatomic point of view, microtubules are also oriented parallel to the axis of the G fibre [6]. Microtubules are heterodimers formed of α- and β-tubulins (TUA and TUB). While Arabidopsis has respectively six and nine genes of TUA and TUB, nine and 20 genes were identified in the poplar genome [87]. These gene families were studied at the expression level: some genes were predominantly expressed in secondary xylem and induced during tension wood formation. Several of these genes were among the most abundant transcripts detected by micro-array or EST analysis during normal or tension wood formation in poplar or Eucalyptus [40,48,51]. In loblolly pine (Pinus taeda), in an association genetics study of wood quality traits, a TUA gene was found to be significantly associated with MFA in early-wood [88]. Finally, in Eucalyptus, the over-expression of a β-tubulin gene altered MFA in fibres in induced somatic sector analysis [89]. These different lines of evidence strongly suggest that microtubules are involved in MFA control in fibre cell walls. Recently, a poplar gene coding for a Microtubule-Associated Protein (PttMAP20) was shown to be strongly up-regulated in developing xylem. The protein could bind microtubules and was the target of the herbicide 2,6-Dichlorobenzonitrile (DCB), which inhibits cellulose synthesis in plants [90]. This protein could be part of the coupling complex between cellulose synthesis protein machinery and cortical microtubules.
4 Conclusion
Although wood formation is a complex developmental process, some aspects are beginning to be understood at the molecular and cellular level, thanks to genomics and transgenic studies. As wood is a heterogeneous tissue, a lot of progress has to be expected through the development of very precise sampling techniques (laser microdissection) or in vivo imaging. This research area will gain a lot by constant comparisons with what is known in Arabidopsis, as this model plant system shows similar processes to trees, such as lignification and cellulose synthesis, while it has its limits for the study of mechanisms underlying heartwood formation or perennial life. The availability of the poplar genome sequence has facilitated comparative studies and shows that some gene families are very much expanded in poplar, suggesting that some families are involved in specific aspects of tree biology.
Acknowledgements
We thank Dr David Lees for English proofreading of the manuscript.


